 |
PDBsum entry 6dv4

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Hydrolase, antiviral protein
|
PDB id
|
|

|

|
6dv4
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |
|
|
 |
 |
 |
 |
Enzyme class 1:
|
 |
E.C.2.7.7.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 2:
|
 |
E.C.2.7.7.49
- RNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
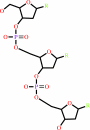 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
E.C.2.7.7.7
- DNA-directed Dna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
DNA(n) + a 2'-deoxyribonucleoside 5'-triphosphate = DNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
 DNA(n)
DNA(n)
|
+
|
2'-deoxyribonucleoside 5'-triphosphate
|
=
|
DNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 4:
|
 |
E.C.3.1.-.-
|
|
 |
 |
 |
 |
 |
Enzyme class 5:
|
 |
E.C.3.1.13.2
- exoribonuclease H.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Exonucleolytic cleavage to 5'-phosphomonoester oligonucleotides in both 5'- to 3'- and 3'- to 5'-directions.
|
 |
 |
 |
 |
 |
Enzyme class 6:
|
 |
E.C.3.1.26.13
- retroviral ribonuclease H.
|
|
 |
 |
 |
 |
 |
Enzyme class 7:
|
 |
E.C.3.4.23.16
- HIV-1 retropepsin.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Specific for a P1 residue that is hydrophobic, and P1' variable, but often Pro.
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
DOI no:
|
Eur J Med Chem
160:171-182
(2018)
|
|
PubMed id:
|
|
|
|

|
| |
|
Design, synthesis, and X-ray studies of potent HIV-1 protease inhibitors incorporating aminothiochromane and aminotetrahydronaphthalene carboxamide derivatives as the P2 ligands.
|
|
A.K.Ghosh,
R.D.Jadhav,
H.Simpson,
S.Kovela,
H.Osswald,
J.Agniswamy,
Y.F.Wang,
S.I.Hattori,
I.T.Weber,
H.Mitsuya.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
We describe the design, synthesis, and biological evaluation of a series of
novel HIV-1 protease inhibitors with carboxamide derivatives as the P2 ligands.
We have specifically designed aminothiochromane and
aminotetrahydronaphthalene-based carboxamide ligands to promote hydrogen bonding
and van der Waals interactions in the active site of HIV-1 protease. Inhibitors
4e and 4j have shown potent enzyme inhibitory and antiviral activity. High
resolution X-ray crystal structures of 4d- and 4k-bound HIV-1 protease revealed
molecular insights into the ligand-binding site interactions.

|

|
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

