 |
PDBsum entry 4hdh
 |
|
|
 |
 |
 |
 |
Enzyme class 1:
|
 |
E.C.2.1.1.56
- mRNA (guanine-N(7))-methyltransferase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L- methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-homocysteine
|
 |
 |
 |
 |
 |
5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L- methionine
S-adenosyl-L- methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
S-adenosyl-L-homocysteine
Bound ligand (Het Group name = )
matches with 46.15% similarity
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 2:
|
 |
E.C.2.1.1.57
- methyltransferase cap1.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA + S-adenosyl-L-methionine = a 5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA + S-adenosyl-L-homocysteine + H+
|
 |
 |
 |
 |
 |
5'-end (N(7)-methyl 5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 S-adenosyl-L-methionine
S-adenosyl-L-methionine
|
=
|
5'-end (N(7)-methyl 5'-triphosphoguanosine)- (2'-O-methyl-ribonucleoside) in mRNA
|
+
|
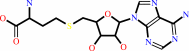 S-adenosyl-L-homocysteine
S-adenosyl-L-homocysteine
|
+
|
H(+)
Bound ligand (Het Group name = )
matches with 46.15% similarity
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
E.C.2.7.7.48
- RNA-directed Rna polymerase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
RNA(n) + a ribonucleoside 5'-triphosphate = RNA(n+1) + diphosphate
|
 |
 |
 |
 |
 |
 RNA(n)
RNA(n)
|
+
|
ribonucleoside 5'-triphosphate
|
=
|
RNA(n+1)
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 4:
|
 |
E.C.3.4.21.91
- flavivirin.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Selective hydrolysis of Xaa-Xaa-|-Xbb bonds in which each of the Xaa can be either Arg or Lys and Xbb can be either Ser or Ala.
|
 |
 |
 |
 |
 |
Enzyme class 5:
|
 |
E.C.3.6.1.15
- nucleoside-triphosphate phosphatase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a ribonucleoside 5'-triphosphate + H2O = a ribonucleoside 5'-diphosphate + phosphate + H+
|
 |
 |
 |
 |
 |
ribonucleoside 5'-triphosphate
|
+
|
H2O
|
=
|
ribonucleoside 5'-diphosphate
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 6:
|
 |
E.C.3.6.4.13
- Rna helicase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
ATP + H2O = ADP + phosphate + H+
|
 |
 |
 |
 |
 |
ATP
Bound ligand (Het Group name = )
corresponds exactly
|
+
|
H2O
|
=
|
 ADP
ADP
|
+
|
 phosphate
phosphate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
DOI no:
|
Nucleic Acids Res
42:2758-2773
(2014)
|
|
PubMed id:
|
|
|
|

|
| |
|
RNA-dependent RNA polymerase of Japanese encephalitis virus binds the initiator nucleotide GTP to form a mechanistically important pre-initiation state.
|
|
P.Surana,
V.Satchidanandam,
D.T.Nair.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
Flaviviral RNA-dependent RNA polymerases (RdRps) initiate replication of the
single-stranded RNA genome in the absence of a primer. The template sequence
5'-CU-3' at the 3'-end of the flaviviral genome is highly conserved.
Surprisingly, flaviviral RdRps require high concentrations of the second
incoming nucleotide GTP to catalyze de novo template-dependent RNA synthesis. We
show that GTP stimulates de novo RNA synthesis by RdRp from Japanese
encephalitis virus (jRdRp) also. Crystal structures of jRdRp complexed with GTP
and ATP provide a basis for specific recognition of GTP. Comparison of the
jRdRpGTP structure with other viral RdRp-GTP structures shows that GTP binds
jRdRp in a novel conformation. Apo-jRdRp structure suggests that the conserved
motif F of jRdRp occupies multiple conformations in absence of GTP. Motif F
becomes ordered on GTP binding and occludes the nucleotide triphosphate entry
tunnel. Mutational analysis of key residues that interact with GTP evinces that
the jRdRpGTP structure represents a novel pre-initiation state. Also, binding
studies show that GTP binding reduces affinity of RdRp for RNA, but the presence
of the catalytic Mn(2+) ion abolishes this inhibition. Collectively, these
observations suggest that the observed pre-initiation state may serve as a
checkpoint to prevent erroneous template-independent RNA synthesis by jRdRp
during initiation.

|

|
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

