Assemblies
Multimeric state:
hetero decamer
Accessible surface area:
96820.16 Å2
Buried surface area:
48398.43 Å2
Dissociation area:
538.74
Å2
Dissociation energy (ΔGdiss):
4.29
kcal/mol
Dissociation entropy (TΔSdiss):
5.92
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-138208
Macromolecules
Chain: A
Length: 826 amino acids
Theoretical weight: 93.78 KDa
Source organism: Homo sapiens
UniProt:
Pfam:
InterPro: Oligosaccharyl transferase, STT3 subunit
Length: 826 amino acids
Theoretical weight: 93.78 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 Q8TCJ2 (Residues: 1-826; Coverage: 100%)
Q8TCJ2 (Residues: 1-826; Coverage: 100%)
Pfam:
InterPro: Oligosaccharyl transferase, STT3 subunit
Chain: B
Length: 37 amino acids
Theoretical weight: 4.2 KDa
Source organism: Homo sapiens
UniProt:
Pfam: Oligosaccaryltransferase
InterPro:
Length: 37 amino acids
Theoretical weight: 4.2 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P0C6T2 (Residues: 1-37; Coverage: 100%)
P0C6T2 (Residues: 1-37; Coverage: 100%)
Pfam: Oligosaccaryltransferase
InterPro:
Chain: C
Length: 79 amino acids
Theoretical weight: 9.08 KDa
Source organism: Homo sapiens
UniProt:
Pfam: Oligosaccharyltransferase subunit 5
InterPro: Oligosaccharyltransferase complex subunit
Length: 79 amino acids
Theoretical weight: 9.08 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P61165 (Residues: 1-79; Coverage: 100%)
P61165 (Residues: 1-79; Coverage: 100%)
Pfam: Oligosaccharyltransferase subunit 5
InterPro: Oligosaccharyltransferase complex subunit
Chain: D
Length: 113 amino acids
Theoretical weight: 12.5 KDa
Source organism: Homo sapiens
UniProt:
Pfam: DAD family
InterPro: DAD/Ost2
Length: 113 amino acids
Theoretical weight: 12.5 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P61803 (Residues: 1-113; Coverage: 100%)
P61803 (Residues: 1-113; Coverage: 100%)
Pfam: DAD family
InterPro: DAD/Ost2
Chain: E
Length: 607 amino acids
Theoretical weight: 68.66 KDa
Source organism: Homo sapiens
UniProt:
Pfam: Ribophorin I
InterPro: Ribophorin I
Length: 607 amino acids
Theoretical weight: 68.66 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P04843 (Residues: 1-607; Coverage: 100%)
P04843 (Residues: 1-607; Coverage: 100%)
Pfam: Ribophorin I
InterPro: Ribophorin I
Chain: F
Length: 631 amino acids
Theoretical weight: 69.35 KDa
Source organism: Homo sapiens
UniProt:
Pfam: Oligosaccharyltransferase subunit Ribophorin II
InterPro: Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit Swp1
Length: 631 amino acids
Theoretical weight: 69.35 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P04844 (Residues: 1-631; Coverage: 100%)
P04844 (Residues: 1-631; Coverage: 100%)
Pfam: Oligosaccharyltransferase subunit Ribophorin II
InterPro: Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit Swp1
Chain: G
Length: 456 amino acids
Theoretical weight: 50.75 KDa
Source organism: Homo sapiens
UniProt:
Pfam: Oligosaccharyltransferase 48 kDa subunit beta
InterPro: Dolichyl-diphosphooligosaccharide--protein glycosyltransferase 48kDa subunit
Length: 456 amino acids
Theoretical weight: 50.75 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P39656 (Residues: 1-456; Coverage: 100%)
P39656 (Residues: 1-456; Coverage: 100%)
Pfam: Oligosaccharyltransferase 48 kDa subunit beta
InterPro: Dolichyl-diphosphooligosaccharide--protein glycosyltransferase 48kDa subunit
Chain: H
Length: 335 amino acids
Theoretical weight: 38.08 KDa
Source organism: Homo sapiens
Expression system: Homo sapiens
UniProt:
Pfam: OST3 / OST6 family, transporter family
InterPro:
Length: 335 amino acids
Theoretical weight: 38.08 KDa
Source organism: Homo sapiens
Expression system: Homo sapiens
UniProt:
- Canonical:
 Q9H0U3 (Residues: 1-335; Coverage: 100%)
Q9H0U3 (Residues: 1-335; Coverage: 100%)
Pfam: OST3 / OST6 family, transporter family
InterPro:
Chain: I
Length: 292 amino acids
Theoretical weight: 32.27 KDa
Source organism: Homo sapiens
UniProt:
Pfam: Malectin domain
InterPro:
Length: 292 amino acids
Theoretical weight: 32.27 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 Q14165 (Residues: 1-292; Coverage: 100%)
Q14165 (Residues: 1-292; Coverage: 100%)
Pfam: Malectin domain
InterPro:



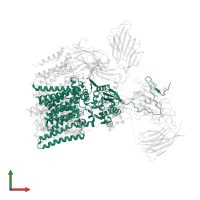
![The deposited structure of PDB entry <span class='highlight'>6s7t</span> coloured by chain, front view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit STT3B</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 4</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Transmembrane protein 258</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit DAD1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 2</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase 48 kDa subunit</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Magnesium transporter protein 1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Malectin</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>PEPTIDE</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)-alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>10 copies of <span class='highlight'>(4R,7R)-4-hydroxy-N,N,N-trimethyl-4,9-dioxo-7-[(undecanoyloxy)methyl]-3,5,8-trioxa-4lambda~5~-phosphadocosan-1-aminium</span>;</li> <li class='image_legend_li'>13 copies of <span class='highlight'>(2~{S},3~{R},4~{R},5~{S},6~{S})-2-(hydroxymethyl)-6-[(1~{S},2~{R},3~{R},4~{R},5'~{S},6~{S},7~{R},8~{S},9~{R},12~{R},13~{R},15~{S},16~{S},18~{R})-5',7,9,13-tetramethyl-3,15-bis(oxidanyl)spiro[5-oxapentacyclo[10.8.0.0^{2,9}.0^{4,8}.0^{13,18}]icosane-6,2'-oxane]-16-yl]oxy-oxane-3,4,5-triol</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>MAGNESIUM ION</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>(2Z,6Z,10Z,14Z,18Z,22Z,26Z)-3,7,11,15,19,23,27,31-octamethyldotriaconta-2,6,10,14,18,22,26,30-octaen-1-yl dihydrogen phosphate</span>.</li></ul> PDB entry 6s7t coloured by chain, front view.](https://www.ebi.ac.uk/pdbe/static/entry/6s7t_deposited_chain_front_image-200x200.png)
![The deposited structure of PDB entry <span class='highlight'>6s7t</span> coloured by chain, side view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit STT3B</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 4</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Transmembrane protein 258</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit DAD1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 2</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase 48 kDa subunit</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Magnesium transporter protein 1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Malectin</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>PEPTIDE</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)-alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>10 copies of <span class='highlight'>(4R,7R)-4-hydroxy-N,N,N-trimethyl-4,9-dioxo-7-[(undecanoyloxy)methyl]-3,5,8-trioxa-4lambda~5~-phosphadocosan-1-aminium</span>;</li> <li class='image_legend_li'>13 copies of <span class='highlight'>(2~{S},3~{R},4~{R},5~{S},6~{S})-2-(hydroxymethyl)-6-[(1~{S},2~{R},3~{R},4~{R},5'~{S},6~{S},7~{R},8~{S},9~{R},12~{R},13~{R},15~{S},16~{S},18~{R})-5',7,9,13-tetramethyl-3,15-bis(oxidanyl)spiro[5-oxapentacyclo[10.8.0.0^{2,9}.0^{4,8}.0^{13,18}]icosane-6,2'-oxane]-16-yl]oxy-oxane-3,4,5-triol</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>MAGNESIUM ION</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>(2Z,6Z,10Z,14Z,18Z,22Z,26Z)-3,7,11,15,19,23,27,31-octamethyldotriaconta-2,6,10,14,18,22,26,30-octaen-1-yl dihydrogen phosphate</span>.</li></ul> PDB entry 6s7t coloured by chain, side view.](https://www.ebi.ac.uk/pdbe/static/entry/6s7t_deposited_chain_side_image-200x200.png)
![The deposited structure of PDB entry <span class='highlight'>6s7t</span> coloured by chain, top view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit STT3B</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 4</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Transmembrane protein 258</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit DAD1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 2</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase 48 kDa subunit</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Magnesium transporter protein 1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Malectin</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>PEPTIDE</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)-alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>10 copies of <span class='highlight'>(4R,7R)-4-hydroxy-N,N,N-trimethyl-4,9-dioxo-7-[(undecanoyloxy)methyl]-3,5,8-trioxa-4lambda~5~-phosphadocosan-1-aminium</span>;</li> <li class='image_legend_li'>13 copies of <span class='highlight'>(2~{S},3~{R},4~{R},5~{S},6~{S})-2-(hydroxymethyl)-6-[(1~{S},2~{R},3~{R},4~{R},5'~{S},6~{S},7~{R},8~{S},9~{R},12~{R},13~{R},15~{S},16~{S},18~{R})-5',7,9,13-tetramethyl-3,15-bis(oxidanyl)spiro[5-oxapentacyclo[10.8.0.0^{2,9}.0^{4,8}.0^{13,18}]icosane-6,2'-oxane]-16-yl]oxy-oxane-3,4,5-triol</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>MAGNESIUM ION</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>(2Z,6Z,10Z,14Z,18Z,22Z,26Z)-3,7,11,15,19,23,27,31-octamethyldotriaconta-2,6,10,14,18,22,26,30-octaen-1-yl dihydrogen phosphate</span>.</li></ul> PDB entry 6s7t coloured by chain, top view.](https://www.ebi.ac.uk/pdbe/static/entry/6s7t_deposited_chain_top_image-200x200.png)
![Hetero decameric assembly 1 of PDB entry <span class='highlight'>6s7t</span> coloured by chemically distinct molecules, front view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit STT3B</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 4</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Transmembrane protein 258</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit DAD1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 2</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase 48 kDa subunit</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Magnesium transporter protein 1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Malectin</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>PEPTIDE</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)-alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>10 copies of <span class='highlight'>(4R,7R)-4-hydroxy-N,N,N-trimethyl-4,9-dioxo-7-[(undecanoyloxy)methyl]-3,5,8-trioxa-4lambda~5~-phosphadocosan-1-aminium</span>;</li> <li class='image_legend_li'>13 copies of <span class='highlight'>(2~{S},3~{R},4~{R},5~{S},6~{S})-2-(hydroxymethyl)-6-[(1~{S},2~{R},3~{R},4~{R},5'~{S},6~{S},7~{R},8~{S},9~{R},12~{R},13~{R},15~{S},16~{S},18~{R})-5',7,9,13-tetramethyl-3,15-bis(oxidanyl)spiro[5-oxapentacyclo[10.8.0.0^{2,9}.0^{4,8}.0^{13,18}]icosane-6,2'-oxane]-16-yl]oxy-oxane-3,4,5-triol</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>MAGNESIUM ION</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>(2Z,6Z,10Z,14Z,18Z,22Z,26Z)-3,7,11,15,19,23,27,31-octamethyldotriaconta-2,6,10,14,18,22,26,30-octaen-1-yl dihydrogen phosphate</span>.</li></ul>} Hetero decameric assembly 1 of PDB entry 6s7t coloured by chemically distinct molecules, front view.](https://www.ebi.ac.uk/pdbe/static/entry/6s7t_assembly_1_chemically_distinct_molecules_front_image-200x200.png)
![Hetero decameric assembly 1 of PDB entry <span class='highlight'>6s7t</span> coloured by chemically distinct molecules, side view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit STT3B</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 4</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Transmembrane protein 258</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit DAD1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 2</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase 48 kDa subunit</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Magnesium transporter protein 1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Malectin</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>PEPTIDE</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)-alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>10 copies of <span class='highlight'>(4R,7R)-4-hydroxy-N,N,N-trimethyl-4,9-dioxo-7-[(undecanoyloxy)methyl]-3,5,8-trioxa-4lambda~5~-phosphadocosan-1-aminium</span>;</li> <li class='image_legend_li'>13 copies of <span class='highlight'>(2~{S},3~{R},4~{R},5~{S},6~{S})-2-(hydroxymethyl)-6-[(1~{S},2~{R},3~{R},4~{R},5'~{S},6~{S},7~{R},8~{S},9~{R},12~{R},13~{R},15~{S},16~{S},18~{R})-5',7,9,13-tetramethyl-3,15-bis(oxidanyl)spiro[5-oxapentacyclo[10.8.0.0^{2,9}.0^{4,8}.0^{13,18}]icosane-6,2'-oxane]-16-yl]oxy-oxane-3,4,5-triol</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>MAGNESIUM ION</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>(2Z,6Z,10Z,14Z,18Z,22Z,26Z)-3,7,11,15,19,23,27,31-octamethyldotriaconta-2,6,10,14,18,22,26,30-octaen-1-yl dihydrogen phosphate</span>.</li></ul>} Hetero decameric assembly 1 of PDB entry 6s7t coloured by chemically distinct molecules, side view.](https://www.ebi.ac.uk/pdbe/static/entry/6s7t_assembly_1_chemically_distinct_molecules_side_image-200x200.png)
![Hetero decameric assembly 1 of PDB entry <span class='highlight'>6s7t</span> coloured by chemically distinct molecules, top view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit STT3B</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 4</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Transmembrane protein 258</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit DAD1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit 2</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Dolichyl-diphosphooligosaccharide--protein glycosyltransferase 48 kDa subunit</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Magnesium transporter protein 1</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Malectin</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>PEPTIDE</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)-alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>alpha-D-mannopyranose-(1-2)-alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-3)-[alpha-D-mannopyranose-(1-6)]alpha-D-mannopyranose-(1-6)]beta-D-mannopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose</span>;</li> <li class='image_legend_li'>10 copies of <span class='highlight'>(4R,7R)-4-hydroxy-N,N,N-trimethyl-4,9-dioxo-7-[(undecanoyloxy)methyl]-3,5,8-trioxa-4lambda~5~-phosphadocosan-1-aminium</span>;</li> <li class='image_legend_li'>13 copies of <span class='highlight'>(2~{S},3~{R},4~{R},5~{S},6~{S})-2-(hydroxymethyl)-6-[(1~{S},2~{R},3~{R},4~{R},5'~{S},6~{S},7~{R},8~{S},9~{R},12~{R},13~{R},15~{S},16~{S},18~{R})-5',7,9,13-tetramethyl-3,15-bis(oxidanyl)spiro[5-oxapentacyclo[10.8.0.0^{2,9}.0^{4,8}.0^{13,18}]icosane-6,2'-oxane]-16-yl]oxy-oxane-3,4,5-triol</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>MAGNESIUM ION</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>(2Z,6Z,10Z,14Z,18Z,22Z,26Z)-3,7,11,15,19,23,27,31-octamethyldotriaconta-2,6,10,14,18,22,26,30-octaen-1-yl dihydrogen phosphate</span>.</li></ul>} Hetero decameric assembly 1 of PDB entry 6s7t coloured by chemically distinct molecules, top view.](https://www.ebi.ac.uk/pdbe/static/entry/6s7t_assembly_1_chemically_distinct_molecules_top_image-200x200.png)