Assemblies
Assembly Name:
Glycosyl hydrolase, family 31
Multimeric state:
monomeric
Accessible surface area:
24500.87 Å2
Buried surface area:
165.03 Å2
Dissociation area:
82.52
Å2
Dissociation energy (ΔGdiss):
-1.02
kcal/mol
Dissociation entropy (TΔSdiss):
0.79
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-107828
Assembly Name:
Glycosyl hydrolase, family 31
Multimeric state:
monomeric
Accessible surface area:
24413.24 Å2
Buried surface area:
0.0 Å2
Dissociation area:
0
Å2
Dissociation energy (ΔGdiss):
0
kcal/mol
Dissociation entropy (TΔSdiss):
0
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-107828
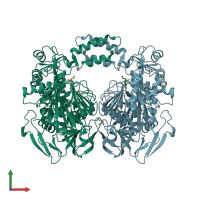
Assembly Name:
Glycosyl hydrolase, family 31
Multimeric state:
homo dimer
Accessible surface area:
43742.33 Å2
Buried surface area:
5336.8 Å2
Dissociation area:
2,551.07
Å2
Dissociation energy (ΔGdiss):
19.55
kcal/mol
Dissociation entropy (TΔSdiss):
15.68
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-107829
Macromolecules
Chains: A, B
Length: 666 amino acids
Theoretical weight: 77.43 KDa
Source organism: Blautia obeum ATCC 29174
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam:
Length: 666 amino acids
Theoretical weight: 77.43 KDa
Source organism: Blautia obeum ATCC 29174
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 A5ZY13 (Residues: 1-663; Coverage: 100%)
A5ZY13 (Residues: 1-663; Coverage: 100%)
Pfam:
- Glycosyl hydrolase 31 N-terminal galactose mutarotase-like domain
- Glycosyl hydrolases family 31 TIM-barrel domain
- Glycosyl hydrolase family 31 C-terminal domain
- Galactose mutarotase-like domain superfamily
- Glycoside hydrolase family 31, N-terminal domain
- Glycoside hydrolase family 31, TIM barrel domain
- Glycoside hydrolase superfamily
- Glycosyl hydrolases family 31, active site