Assemblies
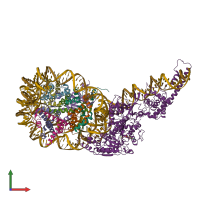
Multimeric state:
hetero undecamer
Accessible surface area:
128511.28 Å2
Buried surface area:
60908.9 Å2
Dissociation area:
4,149.48
Å2
Dissociation energy (ΔGdiss):
59.2
kcal/mol
Dissociation entropy (TΔSdiss):
12.8
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-119088
Macromolecules
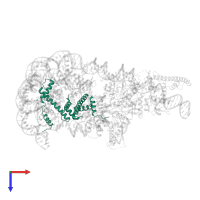
Chains: A, E
Length: 136 amino acids
Theoretical weight: 15.44 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli
UniProt:
InterPro:
Length: 136 amino acids
Theoretical weight: 15.44 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli
UniProt:
- Canonical:
 P84233 (Residues: 1-136; Coverage: 100%)
P84233 (Residues: 1-136; Coverage: 100%)
InterPro:
Chains: B, F
Length: 103 amino acids
Theoretical weight: 11.39 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli
UniProt:
InterPro:
Length: 103 amino acids
Theoretical weight: 11.39 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli
UniProt:
- Canonical:
 P62799 (Residues: 1-103; Coverage: 100%)
P62799 (Residues: 1-103; Coverage: 100%)
InterPro:
Chains: C, G
Length: 130 amino acids
Theoretical weight: 14.11 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli
UniProt:
InterPro:
Length: 130 amino acids
Theoretical weight: 14.11 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli
UniProt:
- Canonical:
 P06897 (Residues: 1-130; Coverage: 100%)
P06897 (Residues: 1-130; Coverage: 100%)
InterPro:
Chain: D
Length: 123 amino acids
Theoretical weight: 13.66 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli
UniProt:
InterPro:
Length: 123 amino acids
Theoretical weight: 13.66 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli
UniProt:
- Canonical:
 P02281 (Residues: 5-126; Coverage: 97%)
P02281 (Residues: 5-126; Coverage: 97%)
InterPro:
Chain: H
Length: 122 amino acids
Theoretical weight: 13.53 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli
UniProt:
InterPro:
Length: 122 amino acids
Theoretical weight: 13.53 KDa
Source organism: Xenopus laevis
Expression system: Escherichia coli
UniProt:
- Canonical:
 P02281 (Residues: 5-125; Coverage: 96%)
P02281 (Residues: 5-125; Coverage: 96%)
InterPro:
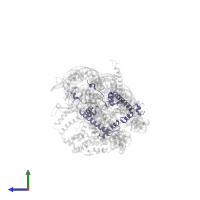
Chain: W
Length: 1468 amino acids
Theoretical weight: 168.5 KDa
Source organism: Saccharomyces cerevisiae S288C
Expression system: Trichoplusia ni
UniProt:
Pfam:
Length: 1468 amino acids
Theoretical weight: 168.5 KDa
Source organism: Saccharomyces cerevisiae S288C
Expression system: Trichoplusia ni
UniProt:
- Canonical:
 P32657 (Residues: 1-1468; Coverage: 100%)
P32657 (Residues: 1-1468; Coverage: 100%)
Pfam:
- Chromo (CHRromatin Organisation MOdifier) domain
- SNF2-related domain
- Helicase conserved C-terminal domain
- Chromodomain helicase DNA-binding domain 1
- Chromodomain-helicase-DNA-binding protein 1-like, C-terminal
- Chromo-like domain superfamily
- Chromo/chromo shadow domain
- Chromo domain
- Chromo domain, conserved site
- P-loop containing nucleoside triphosphate hydrolase
- SNF2-like, N-terminal domain superfamily
- Helicase superfamily 1/2, ATP-binding domain
- SNF2, N-terminal
- Helicase, C-terminal domain-like
- Chromodomain helicase DNA-binding domain
- Homeobox-like domain superfamily
- Chromodomain-helicase-DNA-binding protein 1-like, C-terminal domain