Assemblies
Assembly Name:
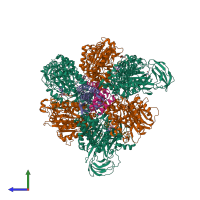
V-type sodium ATPase
Multimeric state:
hetero octamer
Accessible surface area:
114782.08 Å2
Buried surface area:
44920.71 Å2
Dissociation area:
2,260.33
Å2
Dissociation energy (ΔGdiss):
27.57
kcal/mol
Dissociation entropy (TΔSdiss):
12.61
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-155071
Macromolecules
Chains: A, B, C
Length: 600 amino acids
Theoretical weight: 66.35 KDa
Source organism: Enterococcus hirae ATCC 9790
Expression system: Escherichia coli
UniProt:
Pfam:
Length: 600 amino acids
Theoretical weight: 66.35 KDa
Source organism: Enterococcus hirae ATCC 9790
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q08636 (Residues: 1-593; Coverage: 100%)
Q08636 (Residues: 1-593; Coverage: 100%)
Pfam:
- ATP synthase alpha/beta family, beta-barrel domain
- ATPsynthase alpha/beta subunit N-term extension
- ATP synthase alpha/beta family, nucleotide-binding domain
- ATP synthase subunit alpha, N-terminal domain-like superfamily
- V-type ATP synthase catalytic alpha chain
- ATPase, F1/V1/A1 complex, alpha/beta subunit, N-terminal domain superfamily
- ATPase, F1/V1/A1 complex, alpha/beta subunit, N-terminal domain
- P-loop containing nucleoside triphosphate hydrolase
- ATPsynthase alpha/beta subunit, N-terminal extension
- ATPase, F1/V1/A1 complex, alpha/beta subunit, nucleotide-binding domain
- ATPase, alpha/beta subunit, nucleotide-binding domain, active site
- ATPase, F1/V1 complex, beta/alpha subunit, C-terminal
Chains: D, E, F
Length: 465 amino acids
Theoretical weight: 51.72 KDa
Source organism: Enterococcus hirae ATCC 9790
Expression system: Escherichia coli
UniProt:
Pfam:
Length: 465 amino acids
Theoretical weight: 51.72 KDa
Source organism: Enterococcus hirae ATCC 9790
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q08637 (Residues: 1-458; Coverage: 100%)
Q08637 (Residues: 1-458; Coverage: 100%)
Pfam:
- ATP synthase alpha/beta family, beta-barrel domain
- ATP synthase alpha/beta family, nucleotide-binding domain
- ATPase, F1/V1/A1 complex, alpha/beta subunit, N-terminal domain
- P-loop containing nucleoside triphosphate hydrolase
- ATPase, F1/V1/A1 complex, alpha/beta subunit, nucleotide-binding domain
- ATPase, alpha/beta subunit, nucleotide-binding domain, active site
- V-type ATP synthase regulatory subunit B/beta
Chain: G
Length: 217 amino acids
Theoretical weight: 25.12 KDa
Source organism: Enterococcus hirae ATCC 9790
Expression system: Escherichia coli
UniProt:
Pfam: ATP synthase subunit D
InterPro: ATPase, V1 complex, subunit D
Length: 217 amino acids
Theoretical weight: 25.12 KDa
Source organism: Enterococcus hirae ATCC 9790
Expression system: Escherichia coli
UniProt:
- Canonical:
 P43435 (Residues: 1-210; Coverage: 100%)
P43435 (Residues: 1-210; Coverage: 100%)
Pfam: ATP synthase subunit D
InterPro: ATPase, V1 complex, subunit D
Chain: H
Length: 115 amino acids
Theoretical weight: 12.69 KDa
Source organism: Enterococcus hirae ATCC 9790
Expression system: Escherichia coli
UniProt:
Pfam: ATP synthase (F/14-kDa) subunit
InterPro:
Length: 115 amino acids
Theoretical weight: 12.69 KDa
Source organism: Enterococcus hirae ATCC 9790
Expression system: Escherichia coli
UniProt:
- Canonical:
 P43455 (Residues: 1-103; Coverage: 100%)
P43455 (Residues: 1-103; Coverage: 100%)
Pfam: ATP synthase (F/14-kDa) subunit
InterPro:
- ATPase, V1 complex, subunit F superfamily
- ATPase, V1 complex, subunit F, bacterial/archaeal
- ATPase, V1 complex, subunit F