Assemblies
Assembly Name:
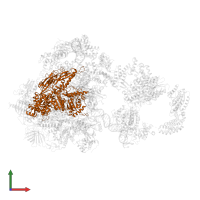
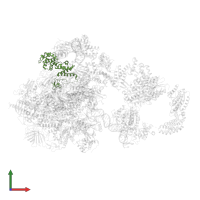
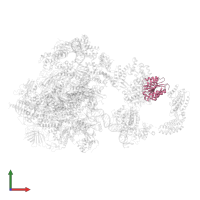
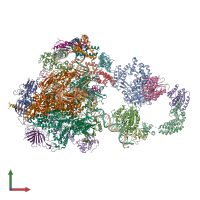
General transcription factor TFIIB-TBP complex and DNA
Multimeric state:
hetero 29-mer
Accessible surface area:
362754.52 Å2
Buried surface area:
114870.39 Å2
Dissociation area:
6,397.88
Å2
Dissociation energy (ΔGdiss):
-14.19
kcal/mol
Dissociation entropy (TΔSdiss):
84.96
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-118815
Macromolecules
Chain: A
Length: 1970 amino acids
Theoretical weight: 217.42 KDa
Source organism: Homo sapiens
UniProt:
Pfam:
Length: 1970 amino acids
Theoretical weight: 217.42 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P24928 (Residues: 1-1970; Coverage: 100%)
P24928 (Residues: 1-1970; Coverage: 100%)
Pfam:
- RNA polymerase Rpb1, domain 1
- RNA polymerase Rpb1, domain 2
- RNA polymerase Rpb1, domain 3
- RNA polymerase Rpb1, domain 4
- RNA polymerase Rpb1, domain 5
- RNA polymerase Rpb1, domain 6
- RNA polymerase Rpb1, domain 7
- RNA polymerase Rpb1 C-terminal repeat
- RNA polymerase Rpb1, clamp domain superfamily
- RNA polymerase Rpb1, domain 1
- DNA-directed RNA polymerase, subunit beta-prime
- RNA polymerase, N-terminal
- RNA polymerase, alpha subunit
- RNA polymerase Rpb1, domain 3
- RNA polymerase Rpb1, domain 3 superfamily
- RNA polymerase Rpb1, funnel domain superfamily
- RNA polymerase Rpb1, domain 4
- RNA polymerase Rpb1, domain 5
- RNA polymerase Rpb1, domain 6
- RNA polymerase Rpb1, domain 7
- RNA polymerase Rpb1, domain 7 superfamily
- RNA polymerase II, heptapeptide repeat, eukaryotic
Chain: B
Length: 1174 amino acids
Theoretical weight: 134.07 KDa
Source organism: Homo sapiens
UniProt:
Pfam:
Length: 1174 amino acids
Theoretical weight: 134.07 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P30876 (Residues: 1-1174; Coverage: 100%)
P30876 (Residues: 1-1174; Coverage: 100%)
Pfam:
- RNA polymerase beta subunit
- RNA polymerase Rpb2, domain 2
- RNA polymerase Rpb2, domain 3
- RNA polymerase Rpb2, domain 4
- RNA polymerase Rpb2, domain 5
- RNA polymerase Rpb2, domain 6
- RNA polymerase Rpb2, domain 7
- DNA-directed RNA polymerase, subunit 2
- RNA polymerase, beta subunit, protrusion
- RNA polymerase Rpb2, domain 2
- RNA polymerase Rpb2, domain 2 superfamily
- RNA polymerase Rpb2, domain 3
- RNA polymerase Rpb2, domain 4
- RNA polymerase Rpb2, domain 5
- DNA-directed RNA polymerase, subunit 2, hybrid-binding domain
- DNA-directed RNA polymerase, subunit 2, hybrid-binding domain superfamily
- RNA polymerase Rpb2, OB-fold
- RNA polymerase, beta subunit, conserved site
- RNA polymerase Rpb2, domain 7
Chain: C
Length: 275 amino acids
Theoretical weight: 31.48 KDa
Source organism: Homo sapiens
UniProt:
Pfam:
InterPro:
Length: 275 amino acids
Theoretical weight: 31.48 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P19387 (Residues: 1-275; Coverage: 100%)
P19387 (Residues: 1-275; Coverage: 100%)
Pfam:
InterPro:
Chain: D
Length: 142 amino acids
Theoretical weight: 16.33 KDa
Source organism: Homo sapiens
UniProt:
Pfam: RNA polymerase Rpb4
InterPro:
Length: 142 amino acids
Theoretical weight: 16.33 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 O15514 (Residues: 1-142; Coverage: 100%)
O15514 (Residues: 1-142; Coverage: 100%)
Pfam: RNA polymerase Rpb4
InterPro:
Chain: E
Length: 210 amino acids
Theoretical weight: 24.58 KDa
Source organism: Homo sapiens
UniProt:
Pfam:
InterPro:
Length: 210 amino acids
Theoretical weight: 24.58 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P19388 (Residues: 1-210; Coverage: 100%)
P19388 (Residues: 1-210; Coverage: 100%)
Pfam:
InterPro:
Chain: F
Length: 127 amino acids
Theoretical weight: 14.49 KDa
Source organism: Homo sapiens
UniProt:
Pfam: RNA polymerase Rpb6
InterPro:
Length: 127 amino acids
Theoretical weight: 14.49 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P61218 (Residues: 1-127; Coverage: 100%)
P61218 (Residues: 1-127; Coverage: 100%)
Pfam: RNA polymerase Rpb6
InterPro:
Chain: G
Length: 172 amino acids
Theoretical weight: 19.31 KDa
Source organism: Homo sapiens
UniProt:
Pfam:
InterPro:
Length: 172 amino acids
Theoretical weight: 19.31 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P62487 (Residues: 1-172; Coverage: 100%)
P62487 (Residues: 1-172; Coverage: 100%)
Pfam:
InterPro:
Chain: H
Length: 150 amino acids
Theoretical weight: 17.16 KDa
Source organism: Homo sapiens
UniProt:
Pfam: RNA polymerase Rpb8
InterPro:
Length: 150 amino acids
Theoretical weight: 17.16 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P52434 (Residues: 1-150; Coverage: 100%)
P52434 (Residues: 1-150; Coverage: 100%)
Pfam: RNA polymerase Rpb8
InterPro:
Chain: I
Length: 125 amino acids
Theoretical weight: 14.54 KDa
Source organism: Homo sapiens
UniProt:
Pfam:
InterPro:
Length: 125 amino acids
Theoretical weight: 14.54 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P36954 (Residues: 1-125; Coverage: 100%)
P36954 (Residues: 1-125; Coverage: 100%)
Pfam:
InterPro:
Chain: J
Length: 67 amino acids
Theoretical weight: 7.66 KDa
Source organism: Homo sapiens
UniProt:
Pfam: RNA polymerases N / 8 kDa subunit
InterPro:
Length: 67 amino acids
Theoretical weight: 7.66 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P62875 (Residues: 1-67; Coverage: 100%)
P62875 (Residues: 1-67; Coverage: 100%)
Pfam: RNA polymerases N / 8 kDa subunit
InterPro:
Chain: K
Length: 117 amino acids
Theoretical weight: 13.31 KDa
Source organism: Homo sapiens
UniProt:
Pfam:
InterPro:
Length: 117 amino acids
Theoretical weight: 13.31 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P52435 (Residues: 1-117; Coverage: 100%)
P52435 (Residues: 1-117; Coverage: 100%)
Pfam:
InterPro:
Chain: L
Length: 58 amino acids
Theoretical weight: 7.02 KDa
Source organism: Homo sapiens
UniProt:
Pfam: DNA directed RNA polymerase, 7 kDa subunit
InterPro:
Length: 58 amino acids
Theoretical weight: 7.02 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P53803 (Residues: 1-58; Coverage: 100%)
P53803 (Residues: 1-58; Coverage: 100%)
Pfam: DNA directed RNA polymerase, 7 kDa subunit
InterPro:
Chain: M
Length: 316 amino acids
Theoretical weight: 34.88 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 316 amino acids
Theoretical weight: 34.88 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q00403 (Residues: 1-316; Coverage: 100%)
Q00403 (Residues: 1-316; Coverage: 100%)
Pfam:
InterPro:
Chain: N
Length: 376 amino acids
Theoretical weight: 41.54 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Transcription factor IIA, alpha/beta subunit
InterPro:
Length: 376 amino acids
Theoretical weight: 41.54 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P52655 (Residues: 1-376; Coverage: 100%)
P52655 (Residues: 1-376; Coverage: 100%)
Pfam: Transcription factor IIA, alpha/beta subunit
InterPro:
Chain: O
Length: 109 amino acids
Theoretical weight: 12.47 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam:
Length: 109 amino acids
Theoretical weight: 12.47 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P52657 (Residues: 1-109; Coverage: 100%)
P52657 (Residues: 1-109; Coverage: 100%)
Pfam:
- Transcription initiation factor IIA, gamma subunit, helical domain
- Transcription initiation factor IIA, gamma subunit
Chain: P
Length: 339 amino acids
Theoretical weight: 37.73 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Transcription factor TFIID (or TATA-binding protein, TBP)
InterPro:
Length: 339 amino acids
Theoretical weight: 37.73 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P20226 (Residues: 1-339; Coverage: 100%)
P20226 (Residues: 1-339; Coverage: 100%)
Pfam: Transcription factor TFIID (or TATA-binding protein, TBP)
InterPro:
Chain: Q
Length: 439 amino acids
Theoretical weight: 49.52 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 439 amino acids
Theoretical weight: 49.52 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P29083 (Residues: 1-439; Coverage: 100%)
P29083 (Residues: 1-439; Coverage: 100%)
Pfam:
InterPro:
Chain: R
Length: 291 amino acids
Theoretical weight: 33.11 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 291 amino acids
Theoretical weight: 33.11 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P29084 (Residues: 1-291; Coverage: 100%)
P29084 (Residues: 1-291; Coverage: 100%)
Pfam:
InterPro:
Chain: S
Length: 517 amino acids
Theoretical weight: 58.34 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Transcription initiation factor IIF, alpha subunit (TFIIF-alpha)
InterPro:
Length: 517 amino acids
Theoretical weight: 58.34 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P35269 (Residues: 1-517; Coverage: 100%)
P35269 (Residues: 1-517; Coverage: 100%)
Pfam: Transcription initiation factor IIF, alpha subunit (TFIIF-alpha)
InterPro:
Chain: T
Length: 249 amino acids
Theoretical weight: 28.43 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 249 amino acids
Theoretical weight: 28.43 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P13984 (Residues: 1-249; Coverage: 100%)
P13984 (Residues: 1-249; Coverage: 100%)
Pfam:
InterPro:
Chain: U
Length: 301 amino acids
Theoretical weight: 34.02 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam:
Length: 301 amino acids
Theoretical weight: 34.02 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P23193 (Residues: 1-301; Coverage: 100%)
P23193 (Residues: 1-301; Coverage: 100%)
Pfam:
- Transcription factor S-II (TFIIS)
- TFIIS helical bundle-like domain
- Transcription factor S-II (TFIIS), central domain
- Zinc finger, TFIIS-type
- Transcription elongation factor, IIS-type
- TFIIS/LEDGF domain superfamily
- Transcription factor IIS, N-terminal
- Transcription elongation factor, TFIIS
- Transcription elongation factor, TFIIS/CRSP70, N-terminal, sub-type
- Transcription elongation factor S-II, central domain superfamily
- Transcription elongation factor S-II, central domain
Chain: V
Length: 782 amino acids
Theoretical weight: 89.4 KDa
Source organism: Homo sapiens
UniProt:
Pfam:
Length: 782 amino acids
Theoretical weight: 89.4 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P19447 (Residues: 1-782; Coverage: 100%)
P19447 (Residues: 1-782; Coverage: 100%)
Pfam:
- Helicase conserved C-terminal domain
- Type III restriction enzyme, res subunit
- ERCC3/RAD25/XPB C-terminal helicase
Chain: W
Length: 760 amino acids
Theoretical weight: 87.02 KDa
Source organism: Homo sapiens
UniProt:
Pfam:
InterPro:
Length: 760 amino acids
Theoretical weight: 87.02 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 P18074 (Residues: 1-760; Coverage: 100%)
P18074 (Residues: 1-760; Coverage: 100%)
Pfam:
InterPro:
- P-loop containing nucleoside triphosphate hydrolase
- Helicase superfamily 1/2, ATP-binding domain, DinG/Rad3-type
- Helicase superfamily 1/2, DinG/Rad3-like
- RAD3/XPD family
- Helicase-like, DEXD box c2 type
- ATP-dependent helicase Rad3/Chl1-like
- RAD3-like helicase, DEAD
- DNA/RNA helicase, ATP-dependent, DEAH-box type, conserved site
- Helical and beta-bridge domain
- ATP-dependent helicase, C-terminal
Chain: 0
Length: 395 amino acids
Theoretical weight: 44.48 KDa
Source organism: Homo sapiens
UniProt:
Pfam:
InterPro:
Length: 395 amino acids
Theoretical weight: 44.48 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 Q13888 (Residues: 1-395; Coverage: 100%)
Q13888 (Residues: 1-395; Coverage: 100%)
Pfam:
InterPro:
Chain: 1
Length: 71 amino acids
Theoretical weight: 8.06 KDa
Source organism: Homo sapiens
UniProt:
Pfam: Transcription factor TFIIH complex subunit Tfb5
InterPro:
Length: 71 amino acids
Theoretical weight: 8.06 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 Q6ZYL4 (Residues: 1-71; Coverage: 100%)
Q6ZYL4 (Residues: 1-71; Coverage: 100%)
Pfam: Transcription factor TFIIH complex subunit Tfb5
InterPro:
Chain: 2
Length: 462 amino acids
Theoretical weight: 52.25 KDa
Source organism: Homo sapiens
UniProt:
Pfam:
InterPro:
Length: 462 amino acids
Theoretical weight: 52.25 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 Q92759 (Residues: 1-462; Coverage: 100%)
Q92759 (Residues: 1-462; Coverage: 100%)
Pfam:
InterPro:
Chain: 3
Length: 308 amino acids
Theoretical weight: 34.42 KDa
Source organism: Homo sapiens
UniProt:
Pfam: Transcription factor Tfb4
InterPro:
Length: 308 amino acids
Theoretical weight: 34.42 KDa
Source organism: Homo sapiens
UniProt:
- Canonical:
 Q13889 (Residues: 1-308; Coverage: 100%)
Q13889 (Residues: 1-308; Coverage: 100%)
Pfam: Transcription factor Tfb4
InterPro: