Assemblies
Assembly Name:
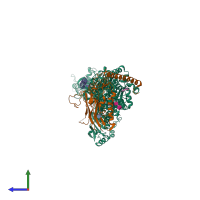
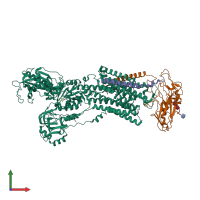
Sodium-Potassium Pump
Multimeric state:
hetero trimer
Accessible surface area:
55910.75 Å2
Buried surface area:
9075.4 Å2
Dissociation area:
1,271.22
Å2
Dissociation energy (ΔGdiss):
10.92
kcal/mol
Dissociation entropy (TΔSdiss):
10.9
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-111292
Macromolecules
Chain: A
Length: 1028 amino acids
Theoretical weight: 113.31 KDa
Source organism: Squalus acanthias
UniProt:
Pfam:
Length: 1028 amino acids
Theoretical weight: 113.31 KDa
Source organism: Squalus acanthias
UniProt:
- Canonical:
 Q4H132 (Residues: 1-1028; Coverage: 100%)
Q4H132 (Residues: 1-1028; Coverage: 100%)
Pfam:
- Cation transporter/ATPase, N-terminus
- E1-E2 ATPase
- Cation transport ATPase (P-type)
- haloacid dehalogenase-like hydrolase
- Cation transporting ATPase, C-terminus
- P-type ATPase subfamily IIC, subunit alpha
- P-type ATPase, transmembrane domain superfamily
- Cation-transporting P-type ATPase, N-terminal
- P-type ATPase
- P-type ATPase, A domain superfamily
- P-type ATPase, haloacid dehalogenase domain
- HAD superfamily
- P-type ATPase, phosphorylation site
- P-type ATPase, cytoplasmic domain N
- HAD-like superfamily
- Cation-transporting P-type ATPase, C-terminal
Chain: B
Length: 305 amino acids
Theoretical weight: 35.18 KDa
Source organism: Squalus acanthias
UniProt:
Pfam: Sodium / potassium ATPase beta chain
InterPro:
Length: 305 amino acids
Theoretical weight: 35.18 KDa
Source organism: Squalus acanthias
UniProt:
- Canonical:
 C4IX13 (Residues: 1-305; Coverage: 100%)
C4IX13 (Residues: 1-305; Coverage: 100%)
Pfam: Sodium / potassium ATPase beta chain
InterPro:
- Sodium/potassium-transporting ATPase subunit beta
- Sodium/potassium-transporting ATPase subunit beta superfamily
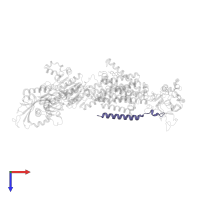
Chain: G
Length: 74 amino acids
Theoretical weight: 8.23 KDa
Source organism: Squalus acanthias
UniProt:
Pfam: ATP1G1/PLM/MAT8 family
InterPro:
Length: 74 amino acids
Theoretical weight: 8.23 KDa
Source organism: Squalus acanthias
UniProt:
- Canonical:
 Q70Q12 (Residues: 21-94; Coverage: 100%)
Q70Q12 (Residues: 21-94; Coverage: 100%)
Pfam: ATP1G1/PLM/MAT8 family
InterPro: