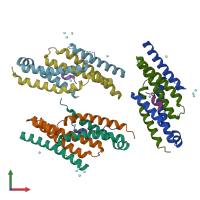
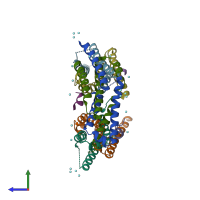
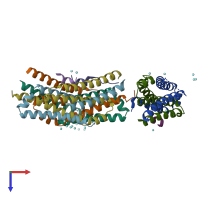
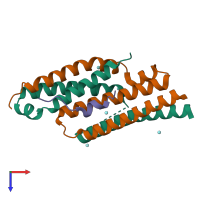
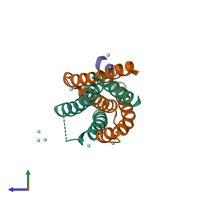
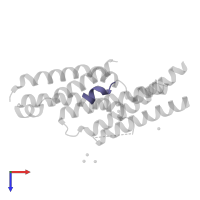
Assemblies
Multimeric state:
hetero trimer
Accessible surface area:
10905.76 Å2
Buried surface area:
8680.86 Å2
Dissociation area:
766.07
Å2
Dissociation energy (ΔGdiss):
6.51
kcal/mol
Dissociation entropy (TΔSdiss):
8.34
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-153725
Multimeric state:
hetero trimer
Accessible surface area:
10628.88 Å2
Buried surface area:
7476.24 Å2
Dissociation area:
0
Å2
Dissociation energy (ΔGdiss):
0
kcal/mol
Dissociation entropy (TΔSdiss):
0
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-153725
Multimeric state:
hetero trimer
Accessible surface area:
10268.38 Å2
Buried surface area:
7188.84 Å2
Dissociation area:
0
Å2
Dissociation energy (ΔGdiss):
0
kcal/mol
Dissociation entropy (TΔSdiss):
0
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-153725
Macromolecules
Chains: A, D, G
Length: 115 amino acids
Theoretical weight: 12.86 KDa
Source organism: Saccharomyces cerevisiae S288C
Expression system: Escherichia coli
UniProt:
Pfam: PMC2NT (NUC016) domain
InterPro: Exosome-associated factor Rrp6, N-terminal
Length: 115 amino acids
Theoretical weight: 12.86 KDa
Source organism: Saccharomyces cerevisiae S288C
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q12149 (Residues: 1-111; Coverage: 15%)
Q12149 (Residues: 1-111; Coverage: 15%)
Pfam: PMC2NT (NUC016) domain
InterPro: Exosome-associated factor Rrp6, N-terminal
Chains: B, E, H
Length: 103 amino acids
Theoretical weight: 12.02 KDa
Source organism: Saccharomyces cerevisiae S288C
Expression system: Escherichia coli
UniProt:
Pfam: Sas10/Utp3/C1D family
InterPro:
Length: 103 amino acids
Theoretical weight: 12.02 KDa
Source organism: Saccharomyces cerevisiae S288C
Expression system: Escherichia coli
UniProt:
- Canonical:
 P38801 (Residues: 1-103; Coverage: 56%)
P38801 (Residues: 1-103; Coverage: 56%)
Pfam: Sas10/Utp3/C1D family
InterPro: