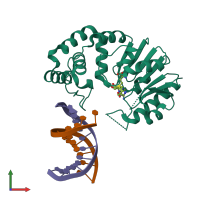Assemblies
Assembly Name:
DNA adenine methylase and DNA
Multimeric state:
hetero trimer
Accessible surface area:
15435.33 Å2
Buried surface area:
1560.59 Å2
Dissociation area:
330.72
Å2
Dissociation energy (ΔGdiss):
-2.61
kcal/mol
Dissociation entropy (TΔSdiss):
10.65
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-118143
Macromolecules
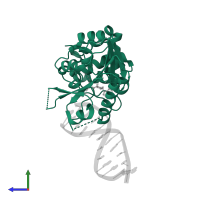
Chain: A
Length: 298 amino acids
Theoretical weight: 34.33 KDa
Source organism: Escherichia coli str. K-12 substr. MDS42
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam: D12 class N6 adenine-specific DNA methyltransferase
InterPro:
Length: 298 amino acids
Theoretical weight: 34.33 KDa
Source organism: Escherichia coli str. K-12 substr. MDS42
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 P0AEE8 (Residues: 1-278; Coverage: 100%)
P0AEE8 (Residues: 1-278; Coverage: 100%)
Pfam: D12 class N6 adenine-specific DNA methyltransferase
InterPro:
- Adenine modification methylase, M.EcoRV-type
- D12 class N6 adenine-specific DNA methyltransferase
- S-adenosyl-L-methionine-dependent methyltransferase superfamily
- Adenine-specific methyltransferase, domain 2
- DNA methylase, N-6 adenine-specific, conserved site
Name:
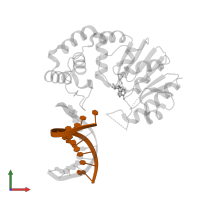
DNA (5'-D(*TP*CP*TP*AP*AP*AP*GP*AP*TP*CP*G)-3')
Representative chains: F
Source organism: synthetic construct [32630]
Expression system: Not provided
Length: 11 nucleotides
Theoretical weight: 3.36 KDa
Representative chains: F
Source organism: synthetic construct [32630]
Expression system: Not provided
Length: 11 nucleotides
Theoretical weight: 3.36 KDa
Name:
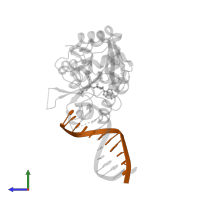
DNA (5'-D(*AP*CP*GP*AP*TP*CP*TP*TP*TP*AP*G)-3')
Representative chains: G
Source organism: synthetic construct [32630]
Expression system: Not provided
Length: 11 nucleotides
Theoretical weight: 3.35 KDa
Representative chains: G
Source organism: synthetic construct [32630]
Expression system: Not provided
Length: 11 nucleotides
Theoretical weight: 3.35 KDa