Assemblies
Assembly Name:
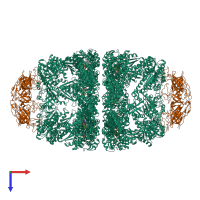
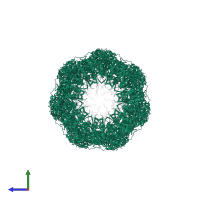
Co-chaperonin GroES and Chaperonin GroEL
Multimeric state:
hetero 28-mer
Accessible surface area:
349802.78 Å2
Buried surface area:
106795.6 Å2
Dissociation area:
1,789.17
Å2
Dissociation energy (ΔGdiss):
4.05
kcal/mol
Dissociation entropy (TΔSdiss):
19.16
kcal/mol
Symmetry number:
3
PDBe Complex ID:
PDB-CPX-176231
Macromolecules
Chains: A, B, C, D, E, F, G, H, I, J, K, L, M, N
Length: 548 amino acids
Theoretical weight: 57.39 KDa
Source organism: Escherichia coli
Expression system: Escherichia coli
UniProt:
Pfam: TCP-1/cpn60 chaperonin family
InterPro:
Length: 548 amino acids
Theoretical weight: 57.39 KDa
Source organism: Escherichia coli
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q548M1 (Residues: 1-548; Coverage: 100%)
Q548M1 (Residues: 1-548; Coverage: 100%)
Pfam: TCP-1/cpn60 chaperonin family
InterPro:
- Chaperonin Cpn60/GroEL
- GroEL-like equatorial domain superfamily
- Chaperonin Cpn60/GroEL/TCP-1 family
- TCP-1-like chaperonin intermediate domain superfamily
- GroEL-like apical domain superfamily
- Chaperonin Cpn60, conserved site
Chains: 1, 2, O, P, Q, R, S, T, U, V, W, X, Y, Z
Length: 97 amino acids
Theoretical weight: 10.4 KDa
Source organism: Escherichia coli
Expression system: Escherichia coli
UniProt:
Pfam: Chaperonin 10 Kd subunit
InterPro:
Length: 97 amino acids
Theoretical weight: 10.4 KDa
Source organism: Escherichia coli
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q7BGE6 (Residues: 1-97; Coverage: 100%)
Q7BGE6 (Residues: 1-97; Coverage: 100%)
Pfam: Chaperonin 10 Kd subunit
InterPro:
- GroES-like superfamily
- GroES chaperonin superfamily
- GroES chaperonin family
- Chaperonin GroES, conserved site