Assemblies
Assembly Name:
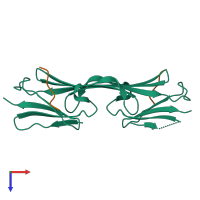
Heat shock protein beta-1
Multimeric state:
hetero tetramer
Accessible surface area:
9913.4 Å2
Buried surface area:
3157.42 Å2
Dissociation area:
743.2
Å2
Dissociation energy (ΔGdiss):
-2.01
kcal/mol
Dissociation entropy (TΔSdiss):
10.93
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-138181
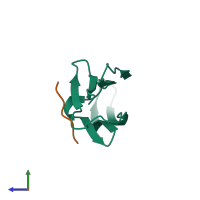
Assembly Name:
Heat shock protein beta-1
Multimeric state:
hetero dimer
Accessible surface area:
5745.9 Å2
Buried surface area:
825.39 Å2
Dissociation area:
412.69
Å2
Dissociation energy (ΔGdiss):
2.67
kcal/mol
Dissociation entropy (TΔSdiss):
6.18
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-138179
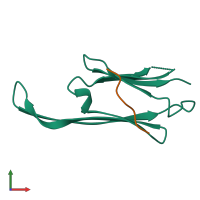
Assembly Name:
Heat shock protein beta-1
Multimeric state:
hetero dimer
Accessible surface area:
5653.91 Å2
Buried surface area:
845.62 Å2
Dissociation area:
422.81
Å2
Dissociation energy (ΔGdiss):
3.2
kcal/mol
Dissociation entropy (TΔSdiss):
6.13
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-138179
Macromolecules
Chains: A, C
Length: 93 amino acids
Theoretical weight: 10.35 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam: Hsp20/alpha crystallin family
InterPro:
Length: 93 amino acids
Theoretical weight: 10.35 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 P04792 (Residues: 84-176; Coverage: 45%)
P04792 (Residues: 84-176; Coverage: 45%)
Pfam: Hsp20/alpha crystallin family
InterPro:
- Alpha crystallin/Small heat shock protein, animal type
- Alpha crystallin/Hsp20 domain
- HSP20-like chaperone
- Heat shock protein beta-1, ACD domain
Chains: B, D
Length: 8 amino acids
Theoretical weight: 919 Da
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
Length: 8 amino acids
Theoretical weight: 919 Da
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
- Canonical:
 P04792 (Residues: 179-186; Coverage: 4%)
P04792 (Residues: 179-186; Coverage: 4%)