Assemblies
Assembly Name:
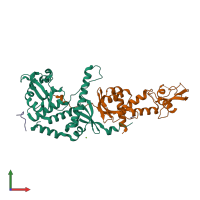
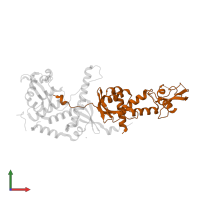
MutLalpha endonuclease complex and Exodeoxyribonuclease 1
Multimeric state:
hetero trimer
Accessible surface area:
24388.63 Å2
Buried surface area:
4756.57 Å2
Dissociation area:
403.18
Å2
Dissociation energy (ΔGdiss):
1.14
kcal/mol
Dissociation entropy (TΔSdiss):
6.65
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-146889
Macromolecules
Chain: A
Length: 288 amino acids
Theoretical weight: 33.24 KDa
Source organism: Saccharomyces cerevisiae S288C
Expression system: Escherichia coli
UniProt:
Pfam: DNA mismatch repair protein Mlh1 C-terminus
InterPro: DNA mismatch repair protein Mlh1, C-terminal
Length: 288 amino acids
Theoretical weight: 33.24 KDa
Source organism: Saccharomyces cerevisiae S288C
Expression system: Escherichia coli
UniProt:
- Canonical:
 P38920 (Residues: 485-769; Coverage: 37%)
P38920 (Residues: 485-769; Coverage: 37%)
Pfam: DNA mismatch repair protein Mlh1 C-terminus
InterPro: DNA mismatch repair protein Mlh1, C-terminal
Chain: B
Length: 240 amino acids
Theoretical weight: 27.95 KDa
Source organism: Saccharomyces cerevisiae S288C
Expression system: Escherichia coli
UniProt:
Pfam: MutL C terminal dimerisation domain
InterPro:
Length: 240 amino acids
Theoretical weight: 27.95 KDa
Source organism: Saccharomyces cerevisiae S288C
Expression system: Escherichia coli
UniProt:
- Canonical:
 P14242 (Residues: 635-873; Coverage: 27%)
P14242 (Residues: 635-873; Coverage: 27%)
Pfam: MutL C terminal dimerisation domain
InterPro: