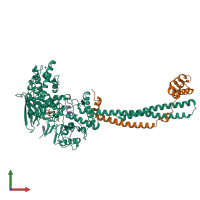
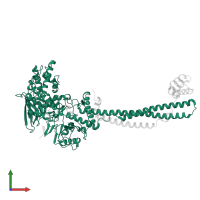
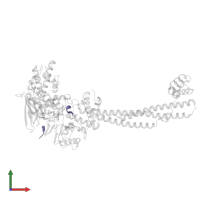
Assemblies
Multimeric state:
hetero tetramer
Accessible surface area:
37716.03 Å2
Buried surface area:
8732.06 Å2
Dissociation area:
331.11
Å2
Dissociation energy (ΔGdiss):
0.89
kcal/mol
Dissociation entropy (TΔSdiss):
5.71
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-129952
Macromolecules
Chain: A
Length: 872 amino acids
Theoretical weight: 94.82 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21
UniProt:
Pfam:
InterPro:
Length: 872 amino acids
Theoretical weight: 94.82 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21
UniProt:
- Canonical:
 O60341 (Residues: 1-173, 174-852; Coverage: 100%)
O60341 (Residues: 1-173, 174-852; Coverage: 100%)
Pfam:
InterPro:
- Lysine-specific histone demethylase
- Homeobox-like domain superfamily
- Winged helix-like DNA-binding domain superfamily
- SWIRM domain
- FAD/NAD(P)-binding domain superfamily
- Amine oxidase
Chain: B
Length: 482 amino acids
Theoretical weight: 53.1 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21
UniProt:
Pfam:
InterPro:
CATH: Methane Monooxygenase Hydroxylase; Chain G, domain 1
Length: 482 amino acids
Theoretical weight: 53.1 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21
UniProt:
- Canonical:
 Q9UKL0 (Residues: 4-485; Coverage: 99%)
Q9UKL0 (Residues: 4-485; Coverage: 99%)
Pfam:
InterPro:
CATH: Methane Monooxygenase Hydroxylase; Chain G, domain 1
Chains: C, D
Length: 6 amino acids
Theoretical weight: 675 Da
Source organism: Homo sapiens
Expression system: Escherichia coli BL21
Length: 6 amino acids
Theoretical weight: 675 Da
Source organism: Homo sapiens
Expression system: Escherichia coli BL21