Assemblies
Assembly Name:
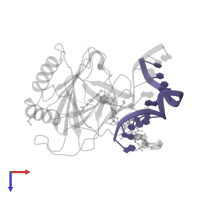
Alpha-ketoglutarate-dependent dioxygenase AlkB and DNA
Multimeric state:
hetero trimer
Accessible surface area:
12550.2 Å2
Buried surface area:
4149.96 Å2
Dissociation area:
1,662.28
Å2
Dissociation energy (ΔGdiss):
4.26
kcal/mol
Dissociation entropy (TΔSdiss):
20.26
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-116632
Macromolecules
Chain: A
Length: 206 amino acids
Theoretical weight: 22.96 KDa
Source organism: Escherichia coli K-12
Expression system: Escherichia coli
UniProt:
InterPro:
Length: 206 amino acids
Theoretical weight: 22.96 KDa
Source organism: Escherichia coli K-12
Expression system: Escherichia coli
UniProt:
- Canonical:
 P05050 (Residues: 12-216; Coverage: 95%)
P05050 (Residues: 12-216; Coverage: 95%)
InterPro:
- Alpha-ketoglutarate-dependent dioxygenase AlkB-like superfamily
- Alpha-ketoglutarate-dependent dioxygenase AlkB-like
- Alkylated DNA repair protein AlkB
- Oxoglutarate/iron-dependent dioxygenase
Name:
DNA (5'-D(*T*AP*GP*GP*TP*AP*AP*(EDA)P*AP*CP*CP*GP*T)-3')
Representative chains: B
Source organism: Escherichia coli K-12 [83333]
Expression system: Not provided
Length: 13 nucleotides
Theoretical weight: 4.08 KDa
Representative chains: B
Source organism: Escherichia coli K-12 [83333]
Expression system: Not provided
Length: 13 nucleotides
Theoretical weight: 4.08 KDa
Name:
DNA (5'-D(*AP*AP*CP*GP*GP*TP*AP*TP*TP*AP*CP*CP*T)-3')
Representative chains: C
Source organism: Escherichia coli K-12 [83333]
Expression system: Not provided
Length: 13 nucleotides
Theoretical weight: 3.95 KDa
Representative chains: C
Source organism: Escherichia coli K-12 [83333]
Expression system: Not provided
Length: 13 nucleotides
Theoretical weight: 3.95 KDa