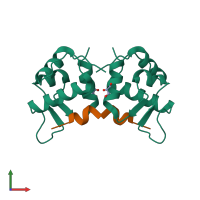Assemblies
Assembly Name:
E3 ubiquitin-protein ligase Mdm2 and peptide
Multimeric state:
hetero tetramer
Accessible surface area:
9801.25 Å2
Buried surface area:
4282.82 Å2
Dissociation area:
906.89
Å2
Dissociation energy (ΔGdiss):
1.5
kcal/mol
Dissociation entropy (TΔSdiss):
11.01
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-163742
Assembly Name:
E3 ubiquitin-protein ligase Mdm2 and peptide
Multimeric state:
hetero tetramer
Accessible surface area:
9603.98 Å2
Buried surface area:
4590.12 Å2
Dissociation area:
1,371.44
Å2
Dissociation energy (ΔGdiss):
12.66
kcal/mol
Dissociation entropy (TΔSdiss):
14.64
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-163742
Macromolecules
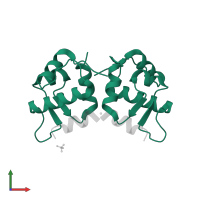
Chains: A, C, E
Length: 85 amino acids
Theoretical weight: 10.04 KDa
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
Pfam: SWIB/MDM2 domain
InterPro:
CATH: SWIB/MDM2 domain
Length: 85 amino acids
Theoretical weight: 10.04 KDa
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
- Canonical:
 Q00987 (Residues: 25-109; Coverage: 17%)
Q00987 (Residues: 25-109; Coverage: 17%)
Pfam: SWIB/MDM2 domain
InterPro:
CATH: SWIB/MDM2 domain
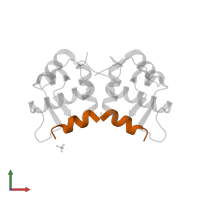
Chains: B, D, F
Length: 12 amino acids
Theoretical weight: 1.52 KDa
Source organism: Homo sapiens
Expression system: Not provided
Length: 12 amino acids
Theoretical weight: 1.52 KDa
Source organism: Homo sapiens
Expression system: Not provided