Assemblies
Assembly Name:
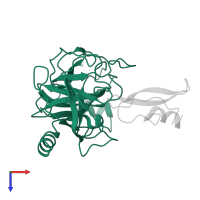
Amyloid-beta protein 40/42 complex and Trypsin-3
Multimeric state:
hetero dimer
Accessible surface area:
12069.13 Å2
Buried surface area:
2681.57 Å2
Dissociation area:
661.88
Å2
Dissociation energy (ΔGdiss):
4.85
kcal/mol
Dissociation entropy (TΔSdiss):
10.3
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-138360
Assembly Name:
Amyloid-beta protein 40/42 complex and Trypsin-3
Multimeric state:
hetero dimer
Accessible surface area:
12046.4 Å2
Buried surface area:
2269.04 Å2
Dissociation area:
658.61
Å2
Dissociation energy (ΔGdiss):
2.29
kcal/mol
Dissociation entropy (TΔSdiss):
10.28
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-138360
Assembly Name:
Amyloid-beta protein 40/42 complex and Trypsin-3
Multimeric state:
hetero dimer
Accessible surface area:
12105.21 Å2
Buried surface area:
1766.75 Å2
Dissociation area:
609.75
Å2
Dissociation energy (ΔGdiss):
2.8
kcal/mol
Dissociation entropy (TΔSdiss):
10.27
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-138360
Assembly Name:
Amyloid-beta protein 40/42 complex and Trypsin-3
Multimeric state:
hetero dimer
Accessible surface area:
11911.53 Å2
Buried surface area:
1507.88 Å2
Dissociation area:
613.12
Å2
Dissociation energy (ΔGdiss):
2.32
kcal/mol
Dissociation entropy (TΔSdiss):
10.26
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-138360
Macromolecules
Chains: A, B, C, D
Length: 224 amino acids
Theoretical weight: 24.26 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Trypsin
InterPro:
Length: 224 amino acids
Theoretical weight: 24.26 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P35030 (Residues: 81-304; Coverage: 74%)
P35030 (Residues: 81-304; Coverage: 74%)
Pfam: Trypsin
InterPro:
- Serine proteases, trypsin domain
- Peptidase S1, PA clan
- Peptidase S1, PA clan, chymotrypsin-like fold
- Peptidase S1A, chymotrypsin family
- Serine proteases, trypsin family, histidine active site
Chains: E, F, G, H
Length: 52 amino acids
Theoretical weight: 5.82 KDa
Source organism: Homo sapiens
Expression system: Komagataella pastoris
UniProt:
Pfam: Kunitz/Bovine pancreatic trypsin inhibitor domain
InterPro:
Length: 52 amino acids
Theoretical weight: 5.82 KDa
Source organism: Homo sapiens
Expression system: Komagataella pastoris
UniProt:
- Canonical:
 P05067 (Residues: 290-341; Coverage: 7%)
P05067 (Residues: 290-341; Coverage: 7%)
Pfam: Kunitz/Bovine pancreatic trypsin inhibitor domain
InterPro:
- Pancreatic trypsin inhibitor Kunitz domain
- Pancreatic trypsin inhibitor Kunitz domain superfamily
- Proteinase inhibitor I2, Kunitz, conserved site