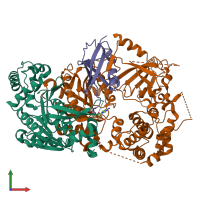
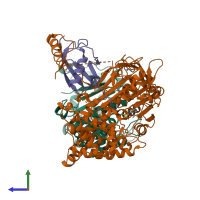
Assemblies
Multimeric state:
hetero trimer
Accessible surface area:
40381.17 Å2
Buried surface area:
9279.91 Å2
Dissociation area:
2,047.97
Å2
Dissociation energy (ΔGdiss):
19.69
kcal/mol
Dissociation entropy (TΔSdiss):
12.15
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-159021
Macromolecules
Chain: A
Length: 346 amino acids
Theoretical weight: 38.5 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: ThiF family
InterPro:
Length: 346 amino acids
Theoretical weight: 38.5 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q9UBE0 (Residues: 1-346; Coverage: 100%)
Q9UBE0 (Residues: 1-346; Coverage: 100%)
Pfam: ThiF family
InterPro:
- Ubiquitin-activating enzyme
- ThiF/MoeB/HesA family
- THIF-type NAD/FAD binding fold
- Ubiquitin/SUMO-activating enzyme E1-like
Chain: B
Length: 660 amino acids
Theoretical weight: 73.5 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam:
Length: 660 amino acids
Theoretical weight: 73.5 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q9UBT2 (Residues: 1-640; Coverage: 100%)
Q9UBT2 (Residues: 1-640; Coverage: 100%)
Pfam:
- ThiF family
- Ubiquitin/SUMO-activating enzyme ubiquitin-like domain
- SUMO-activating enzyme subunit 2 C-terminus
- Ubiquitin-activating enzyme
- ThiF/MoeB/HesA family
- THIF-type NAD/FAD binding fold
- 6-pyruvoyl tetrahydropterin synthase/QueD superfamily
- SUMO-activating enzyme subunit Uba2
- Ubiquitin-activating enzyme E1, inactive adenylation domain, subdomain 1
- Ubiquitin activating enzyme, alpha domain superfamily
- Ubiquitin-activating enzyme E1, Cys active site
- Ubiquitin-activating enzyme E1, conserved site
- Ubiquitin/SUMO-activating enzyme ubiquitin-like domain
- SUMO-activating enzyme subunit 2, C-terminal domain
Chain: D
Length: 97 amino acids
Theoretical weight: 11.15 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Ubiquitin-2 like Rad60 SUMO-like
InterPro:
Length: 97 amino acids
Theoretical weight: 11.15 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P63165 (Residues: 1-97; Coverage: 96%)
P63165 (Residues: 1-97; Coverage: 96%)
Pfam: Ubiquitin-2 like Rad60 SUMO-like
InterPro:
- Ubiquitin-like domain superfamily
- Ubiquitin-like domain
- Small ubiquitin-related modifier 1, Ubl domain
- Rad60/SUMO-like domain