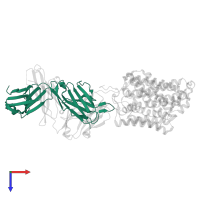
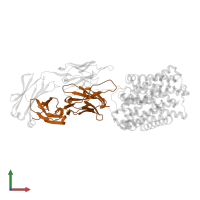
Assemblies
Multimeric state:
hetero trimer
Accessible surface area:
34903.59 Å2
Buried surface area:
5356.09 Å2
Dissociation area:
921.89
Å2
Dissociation energy (ΔGdiss):
1.25
kcal/mol
Dissociation entropy (TΔSdiss):
13.75
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-212286
Macromolecules
Chain: L
Length: 220 amino acids
Theoretical weight: 24.28 KDa
Source organism: Mus musculus
Expression system: Mus musculus
Length: 220 amino acids
Theoretical weight: 24.28 KDa
Source organism: Mus musculus
Expression system: Mus musculus
- Immunoglobulin-like domain
- Immunoglobulin-like fold
- Immunoglobulin-like domain superfamily
- Immunoglobulin subtype
- Immunoglobulin V-set domain
- Immunoglobulin C1-set
- Immunoglobulin/major histocompatibility complex, conserved site
Chain: H
Length: 223 amino acids
Theoretical weight: 23.98 KDa
Source organism: Mus musculus
Expression system: Mus musculus
Length: 223 amino acids
Theoretical weight: 23.98 KDa
Source organism: Mus musculus
Expression system: Mus musculus
- Immunoglobulin-like domain
- Immunoglobulin-like fold
- Immunoglobulin-like domain superfamily
- Immunoglobulin subtype
- Immunoglobulin V-set domain
- Immunoglobulin C1-set
Chain: C
Length: 444 amino acids
Theoretical weight: 48.33 KDa
Source organism: Methanocaldococcus jannaschii
Expression system: Escherichia coli
UniProt:
Pfam: Amino acid permease
InterPro: Amino acid/polyamine transporter I
CATH: Amino acid/polyamine transporter I
Length: 444 amino acids
Theoretical weight: 48.33 KDa
Source organism: Methanocaldococcus jannaschii
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q58026 (Residues: 1-435; Coverage: 100%)
Q58026 (Residues: 1-435; Coverage: 100%)
Pfam: Amino acid permease
InterPro: Amino acid/polyamine transporter I
CATH: Amino acid/polyamine transporter I