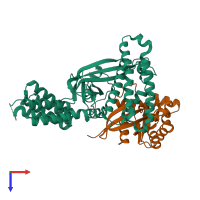
Assemblies
Multimeric state:
hetero dimer
Accessible surface area:
28697.52 Å2
Buried surface area:
3722.82 Å2
Dissociation area:
1,861.41
Å2
Dissociation energy (ΔGdiss):
21.88
kcal/mol
Dissociation entropy (TΔSdiss):
13.18
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-158908
Multimeric state:
hetero dimer
Accessible surface area:
28655.21 Å2
Buried surface area:
3709.21 Å2
Dissociation area:
1,854.6
Å2
Dissociation energy (ΔGdiss):
20.43
kcal/mol
Dissociation entropy (TΔSdiss):
13.15
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-158908
Macromolecules
Chains: A, B
Length: 436 amino acids
Theoretical weight: 51.65 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 436 amino acids
Theoretical weight: 51.65 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q92608 (Residues: 1192-1622; Coverage: 24%)
Q92608 (Residues: 1192-1622; Coverage: 24%)
Pfam:
InterPro:
- Dedicator of cytokinesis, C-terminal, lobe A
- Dedicator of cytokinesis
- Dedicator of cytokinesis protein 2, DHR2 domain
- DOCKER, Lobe A
- DOCKER domain
- DOCKER, Lobe B
- Dedicator of cytokinesis, C-terminal, lobe C
- DOCKER, Lobe C
Chains: C, D
Length: 196 amino acids
Theoretical weight: 21.86 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Ras family
InterPro:
Length: 196 amino acids
Theoretical weight: 21.86 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P63000 (Residues: 1-177; Coverage: 92%)
P63000 (Residues: 1-177; Coverage: 92%)
Pfam: Ras family
InterPro:
- Small GTPase
- P-loop containing nucleoside triphosphate hydrolase
- Small GTP-binding protein domain
- Small GTPase Rho