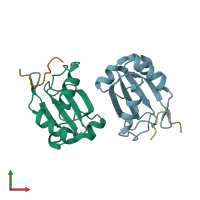
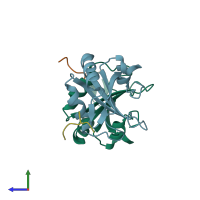
Assemblies
Assembly Name:
Splicing factor 45 and Splicing factor 3B subunit 1
Multimeric state:
hetero dimer
Accessible surface area:
6084.83 Å2
Buried surface area:
1044.0 Å2
Dissociation area:
522
Å2
Dissociation energy (ΔGdiss):
2.54
kcal/mol
Dissociation entropy (TΔSdiss):
7.49
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-131048
Assembly Name:
Splicing factor 45 and Splicing factor 3B subunit 1
Multimeric state:
hetero dimer
Accessible surface area:
5907.11 Å2
Buried surface area:
930.5 Å2
Dissociation area:
465.25
Å2
Dissociation energy (ΔGdiss):
1.44
kcal/mol
Dissociation entropy (TΔSdiss):
6.9
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-131048
Assembly Name:
Splicing factor 45 and Splicing factor 3B subunit 1
Multimeric state:
hetero tetramer
Accessible surface area:
11505.87 Å2
Buried surface area:
2460.56 Å2
Dissociation area:
243.03
Å2
Dissociation energy (ΔGdiss):
-3.9
kcal/mol
Dissociation entropy (TΔSdiss):
10.92
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-131050
Macromolecules
Chains: A, B
Length: 105 amino acids
Theoretical weight: 11.69 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: RNA recognition motif
InterPro:
Length: 105 amino acids
Theoretical weight: 11.69 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q96I25 (Residues: 301-401; Coverage: 25%)
Q96I25 (Residues: 301-401; Coverage: 25%)
Pfam: RNA recognition motif
InterPro:
- Splicing factor 45
- SPF45, RNA recognition motif
- Nucleotide-binding alpha-beta plait domain superfamily
- RNA recognition motif domain, eukaryote
- RNA-binding domain superfamily
- RNA recognition motif domain
Chains: C, D
Length: 10 amino acids
Theoretical weight: 1.31 KDa
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
Length: 10 amino acids
Theoretical weight: 1.31 KDa
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
- Canonical:
 O75533 (Residues: 333-342; Coverage: 1%)
O75533 (Residues: 333-342; Coverage: 1%)