Assemblies
Assembly Name:
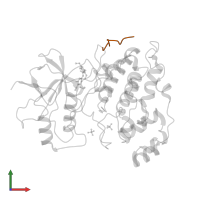
Mitogen-activated protein kinase 8 and C-Jun-amino-terminal kinase-interacting protein 1
Multimeric state:
hetero dimer
Accessible surface area:
17114.63 Å2
Buried surface area:
2359.65 Å2
Dissociation area:
590.86
Å2
Dissociation energy (ΔGdiss):
1.61
kcal/mol
Dissociation entropy (TΔSdiss):
7.71
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-155386
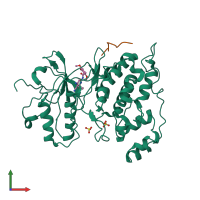
Assembly Name:
Mitogen-activated protein kinase 8 and C-Jun-amino-terminal kinase-interacting protein 1
Multimeric state:
hetero dimer
Accessible surface area:
16952.43 Å2
Buried surface area:
2397.65 Å2
Dissociation area:
567.16
Å2
Dissociation energy (ΔGdiss):
3.83
kcal/mol
Dissociation entropy (TΔSdiss):
7.38
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-155386
Assembly Name:
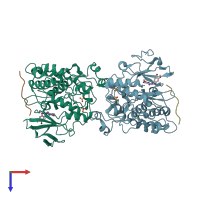
Mitogen-activated protein kinase 8 and C-Jun-amino-terminal kinase-interacting protein 1
Multimeric state:
hetero tetramer
Accessible surface area:
32483.63 Å2
Buried surface area:
6340.72 Å2
Dissociation area:
590.86
Å2
Dissociation energy (ΔGdiss):
1.59
kcal/mol
Dissociation entropy (TΔSdiss):
7.74
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-155389
Macromolecules
Chains: A, B
Length: 370 amino acids
Theoretical weight: 42.92 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Protein kinase domain
InterPro:
Length: 370 amino acids
Theoretical weight: 42.92 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P45983 (Residues: 1-364; Coverage: 85%)
P45983 (Residues: 1-364; Coverage: 85%)
Pfam: Protein kinase domain
InterPro:
- Protein kinase-like domain superfamily
- Protein kinase domain
- Mitogen-activated protein (MAP) kinase, conserved site
- Mitogen-activated protein (MAP) kinase, JNK
- Serine/threonine-protein kinase, active site