Assemblies
Multimeric state:
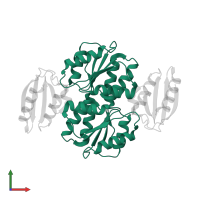
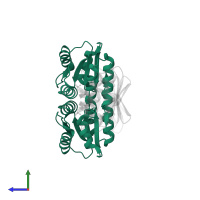
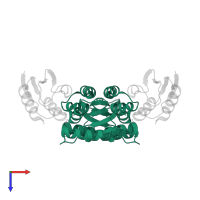
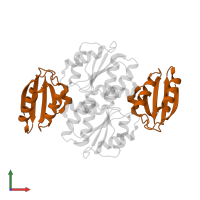
hetero tetramer
Accessible surface area:
17331.58 Å2
Buried surface area:
6852.36 Å2
Dissociation area:
1,606.14
Å2
Dissociation energy (ΔGdiss):
-11.21
kcal/mol
Dissociation entropy (TΔSdiss):
22.58
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-141950
Macromolecules
Chains: A, B
Length: 136 amino acids
Theoretical weight: 14.76 KDa
Source organism: Escherichia coli
Expression system: Escherichia coli
UniProt:
Pfam: PTS system fructose IIA component
InterPro:
SCOP: EIIA-man component-like
Length: 136 amino acids
Theoretical weight: 14.76 KDa
Source organism: Escherichia coli
Expression system: Escherichia coli
UniProt:
- Canonical:
 P69797 (Residues: 1-133; Coverage: 41%)
P69797 (Residues: 1-133; Coverage: 41%)
Pfam: PTS system fructose IIA component
InterPro:
- Phosphotransferase system, mannose-type IIA component
- Phosphotransferase system, mannose family IIA component
- Phosphotransferase system, mannose-type IIA component superfamily
- PTS system mannose/sorbose specific IIA subunit
SCOP: EIIA-man component-like
Chains: C, D
Length: 85 amino acids
Theoretical weight: 9.13 KDa
Source organism: Escherichia coli
Expression system: Escherichia coli
UniProt:
Pfam: PTS HPr component phosphorylation site
InterPro:
SCOP: HPr-like
Length: 85 amino acids
Theoretical weight: 9.13 KDa
Source organism: Escherichia coli
Expression system: Escherichia coli
UniProt:
- Canonical:
 P0AA04 (Residues: 1-85; Coverage: 100%)
P0AA04 (Residues: 1-85; Coverage: 100%)
Pfam: PTS HPr component phosphorylation site
InterPro:
- Phosphocarrier protein HPr-like
- HPr-like superfamily
- Phosphotransferase system, HPr histidine phosphorylation site
- Phosphotransferase system, HPr serine phosphorylation site
SCOP: HPr-like