Assemblies
Assembly Name:
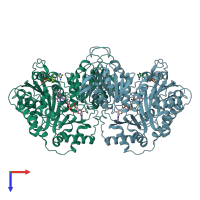
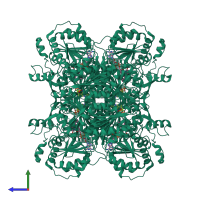
Pyruvate oxidase
Multimeric state:
homo tetramer
Accessible surface area:
70665.69 Å2
Buried surface area:
32454.45 Å2
Dissociation area:
4,264.72
Å2
Dissociation energy (ΔGdiss):
41.82
kcal/mol
Dissociation entropy (TΔSdiss):
16.99
kcal/mol
Symmetry number:
4
PDBe Complex ID:
PDB-CPX-153340
Macromolecules
Chains: A, B
Length: 585 amino acids
Theoretical weight: 64.16 KDa
Source organism: Lactiplantibacillus plantarum
Expression system: Not provided
UniProt:
Pfam:
SCOP:
Length: 585 amino acids
Theoretical weight: 64.16 KDa
Source organism: Lactiplantibacillus plantarum
Expression system: Not provided
UniProt:
- Canonical:
 P37063 (Residues: 9-593; Coverage: 97%)
P37063 (Residues: 9-593; Coverage: 97%)
Pfam:
- Thiamine pyrophosphate enzyme, N-terminal TPP binding domain
- Thiamine pyrophosphate enzyme, central domain
- Thiamine pyrophosphate enzyme, C-terminal TPP binding domain
- Thiamin diphosphate-binding fold
- Pyruvate oxidase
- Pyruvate oxidase POXB-like, pyrimidine-binding domain
- Pyruvate oxidase/Pyruvate dehydrogenase [ubiquinone]-like
- Thiamine pyrophosphate enzyme, N-terminal TPP-binding domain
- DHS-like NAD/FAD-binding domain superfamily
- Thiamine pyrophosphate enzyme, central domain
- Pyruvate oxidase POXB-like, PP-binding domain
- Thiamine pyrophosphate enzyme, TPP-binding
- TPP-binding enzyme, conserved site
SCOP: