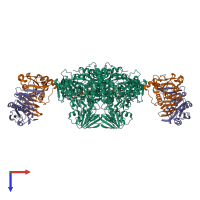
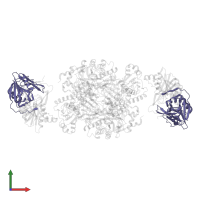
Assemblies
Multimeric state:
hetero hexamer
Accessible surface area:
76441.14 Å2
Buried surface area:
27992.05 Å2
Dissociation area:
998.55
Å2
Dissociation energy (ΔGdiss):
-13.75
kcal/mol
Dissociation entropy (TΔSdiss):
29.22
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-147610
Macromolecules
Chains: A, B
Length: 729 amino acids
Theoretical weight: 81.61 KDa
Source organism: Methylophilus methylotrophus
UniProt:
Pfam:
InterPro:
SCOP:
Length: 729 amino acids
Theoretical weight: 81.61 KDa
Source organism: Methylophilus methylotrophus
UniProt:
- Canonical:
 P16099 (Residues: 2-730; Coverage: 100%)
P16099 (Residues: 2-730; Coverage: 100%)
Pfam:
InterPro:
- Aldolase-type TIM barrel
- Trimethylamine dehydrogenase/Dimethylamine dehydrogenase, FMN-binding domain
- NADH:flavin oxidoreductase/NADH oxidase, N-terminal
- FAD/NAD(P)-binding domain
- FAD/NAD(P)-binding domain superfamily
SCOP:
Chains: C, E
Length: 264 amino acids
Theoretical weight: 28.93 KDa
Source organism: Methylophilus methylotrophus
Expression system: Escherichia coli
UniProt:
Pfam: Electron transfer flavoprotein domain
InterPro:
SCOP: ETFP subunits
Length: 264 amino acids
Theoretical weight: 28.93 KDa
Source organism: Methylophilus methylotrophus
Expression system: Escherichia coli
UniProt:
- Canonical:
 P53570 (Residues: 1-264; Coverage: 100%)
P53570 (Residues: 1-264; Coverage: 100%)
Pfam: Electron transfer flavoprotein domain
InterPro:
- Electron transfer flavoprotein, beta subunit
- Rossmann-like alpha/beta/alpha sandwich fold
- Electron transfer flavoprotein, beta subunit, N-terminal
- Electron transfer flavoprotein, alpha/beta-subunit, N-terminal
- Electron transfer flavoprotein, beta-subunit, conserved site
SCOP: ETFP subunits
Chains: D, F
Length: 320 amino acids
Theoretical weight: 33.62 KDa
Source organism: Methylophilus methylotrophus
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
SCOP: ETFP subunits
Length: 320 amino acids
Theoretical weight: 33.62 KDa
Source organism: Methylophilus methylotrophus
Expression system: Escherichia coli
UniProt:
- Canonical:
 P53571 (Residues: 2-321; Coverage: 100%)
P53571 (Residues: 2-321; Coverage: 100%)
Pfam:
InterPro:
- Rossmann-like alpha/beta/alpha sandwich fold
- Electron transfer flavoprotein, alpha/beta-subunit, N-terminal
- Electron transfer flavoprotein alpha subunit/FixB
- Electron transfer flavoprotein, alpha subunit, N-terminal
- DHS-like NAD/FAD-binding domain superfamily
- Electron transfer flavoprotein, alpha subunit, C-terminal
- Electron transfer flavoprotein subunit alpha, conserved site
SCOP: ETFP subunits