Assemblies
Assembly Name:
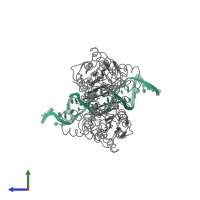
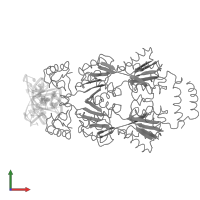
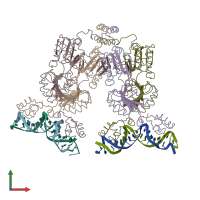
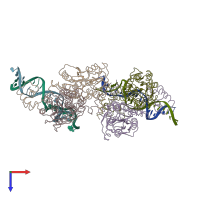
Lactose operon repressor and DNA
Multimeric state:
hetero tetramer
Accessible surface area:
7901.92 Å2
Buried surface area:
1881.74 Å2
Dissociation area:
940.87
Å2
Dissociation energy (ΔGdiss):
16.87
kcal/mol
Dissociation entropy (TΔSdiss):
10.89
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-114128
Assembly Name:
Lactose operon repressor and DNA
Multimeric state:
hetero tetramer
Accessible surface area:
7592.92 Å2
Buried surface area:
2281.31 Å2
Dissociation area:
1,140.66
Å2
Dissociation energy (ΔGdiss):
16.57
kcal/mol
Dissociation entropy (TΔSdiss):
10.9
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-114128
Macromolecules
Chains: A, B, C, D
Length: 360 amino acids
Theoretical weight: 38.66 KDa
Source organism: Escherichia coli
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 360 amino acids
Theoretical weight: 38.66 KDa
Source organism: Escherichia coli
Expression system: Escherichia coli
UniProt:
- Canonical:
 P03023 (Residues: 1-360; Coverage: 100%)
P03023 (Residues: 1-360; Coverage: 100%)
Pfam:
InterPro:
- Lambda repressor-like, DNA-binding domain superfamily
- LacI-type HTH domain
- Periplasmic binding protein-like I
- Transcriptional regulator LacI/GalR-like, sensor domain
Name:
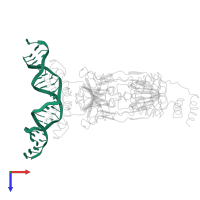
DNA (5'-D(*GP*AP*AP*TP*TP*GP*TP*GP*AP*GP*CP*GP*CP*TP*CP*AP*CP*AP*AP*TP*T)-3')
Representative chains: E, F, G, H
Source organism: Escherichia coli [562]
Expression system: Not provided
Length: 21 nucleotides
Theoretical weight: 6.46 KDa
Representative chains: E, F, G, H
Source organism: Escherichia coli [562]
Expression system: Not provided
Length: 21 nucleotides
Theoretical weight: 6.46 KDa