Assemblies
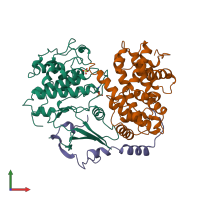
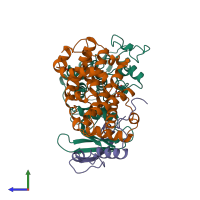
Multimeric state:
hetero trimer
Accessible surface area:
24408.38 Å2
Buried surface area:
9192.86 Å2
Dissociation area:
2,794.99
Å2
Dissociation energy (ΔGdiss):
36.33
kcal/mol
Dissociation entropy (TΔSdiss):
12.62
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-148888
Macromolecules
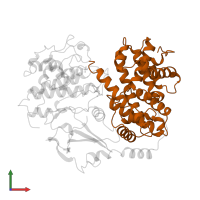
Chain: A
Length: 298 amino acids
Theoretical weight: 34.06 KDa
Source organism: Homo sapiens
Expression system: Spodoptera frugiperda
UniProt:
Pfam: Protein kinase domain
InterPro:
SCOP: Protein kinases, catalytic subunit
Length: 298 amino acids
Theoretical weight: 34.06 KDa
Source organism: Homo sapiens
Expression system: Spodoptera frugiperda
UniProt:
- Canonical:
 P24941 (Residues: 1-298; Coverage: 100%)
P24941 (Residues: 1-298; Coverage: 100%)
Pfam: Protein kinase domain
InterPro:
- Protein kinase-like domain superfamily
- Protein kinase domain
- Protein kinase, ATP binding site
- Serine/threonine-protein kinase, active site
SCOP: Protein kinases, catalytic subunit
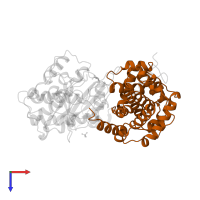
Chain: B
Length: 260 amino acids
Theoretical weight: 29.87 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
SCOP: Cyclin
Length: 260 amino acids
Theoretical weight: 29.87 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P20248 (Residues: 173-432; Coverage: 60%)
P20248 (Residues: 173-432; Coverage: 60%)
Pfam:
InterPro:
- Cyclin A/B-like
- Cyclin-like superfamily
- Cyclin
- Cyclin, N-terminal
- Cyclin-like domain
- Cyclin, C-terminal domain
SCOP: Cyclin
Chain: C
Length: 84 amino acids
Theoretical weight: 9.97 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Cyclin-dependent kinase inhibitor
InterPro:
CATH: p27
SCOP: P27(KIP1) fragment 22-106
Length: 84 amino acids
Theoretical weight: 9.97 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P46527 (Residues: 23-106; Coverage: 42%)
P46527 (Residues: 23-106; Coverage: 42%)
Pfam: Cyclin-dependent kinase inhibitor
InterPro:
CATH: p27
SCOP: P27(KIP1) fragment 22-106