Assemblies
Assembly Name:
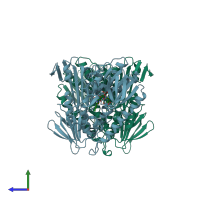
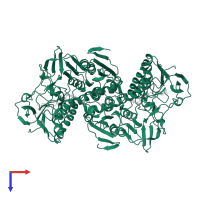
Trypanothione reductase
Multimeric state:
homo dimer
Accessible surface area:
35292.71 Å2
Buried surface area:
9446.91 Å2
Dissociation area:
3,276.24
Å2
Dissociation energy (ΔGdiss):
60.88
kcal/mol
Dissociation entropy (TΔSdiss):
15.22
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-153807
Macromolecules
Chains: A, B
Length: 490 amino acids
Theoretical weight: 53.18 KDa
Source organism: Crithidia fasciculata
Expression system: Escherichia coli
UniProt:
Pfam:
SCOP:
Length: 490 amino acids
Theoretical weight: 53.18 KDa
Source organism: Crithidia fasciculata
Expression system: Escherichia coli
UniProt:
- Canonical:
 P39040 (Residues: 2-491; Coverage: 100%)
P39040 (Residues: 2-491; Coverage: 100%)
Pfam:
- Pyridine nucleotide-disulphide oxidoreductase
- Pyridine nucleotide-disulphide oxidoreductase, dimerisation domain
- Pyridine nucleotide-disulphide oxidoreductase, class I
- Trypanothione reductase
- FAD/NAD(P)-binding domain
- FAD/NAD(P)-binding domain superfamily
- Glutathione reductase/thioredoxin reductase-like
- Pyridine nucleotide-disulphide oxidoreductase, class I, active site
- FAD/NAD-linked reductase, dimerisation domain superfamily
- Pyridine nucleotide-disulphide oxidoreductase, dimerisation domain
SCOP: