EMD-20060
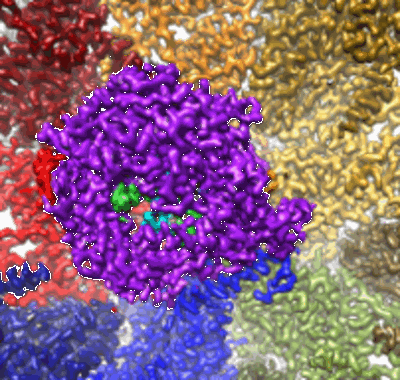
In situ structure of Rotavirus RNA-dependent RNA polymerase at transcript-elongated state
EMD-20060
Single-particle3.6 Å
 Deposition: 03/04/2019
Deposition: 03/04/2019Map released: 22/05/2019
Last modified: 20/03/2024
Name: Rotavirus A
Summary:
Name: RNA (5'-R(P*UP*AP*UP*AP*UP*AP*UP*AP*UP*AP*UP*AP*UP*AP*UP*AP*UP*A)-3')
Number of copies: 1
Number of copies: 1
Natural source
Organism: Rotavirus A
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 5 kDa | - |
Name: RNA (5'-R(P*AP*UP*AP*UP*AP*UP*AP*UP*AP*UP*AP*UP*AP*UP*AP*UP*A)-3')
Number of copies: 1
Number of copies: 1
Natural source
Organism: Rotavirus A
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 5 kDa | - |
Name: RNA-dependent RNA polymerase of rotavirus A
EC number: 2.7.7.48
Number of copies: 1
UniProtKB PDBe-KB
EC number: 2.7.7.48
Number of copies: 1
UniProtKB PDBe-KB
Domains
Gene Ontology [6]
Biological process:
viral genome replication DNA-templated transcription
Molecular function:
nucleotide binding RNA-dependent RNA polymerase activity RNA binding
Cellular location:
virion component
viral genome replication DNA-templated transcription
Molecular function:
nucleotide binding RNA-dependent RNA polymerase activity RNA binding
Cellular location:
virion component
Natural source
Organism: Rotavirus A
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 125 kDa | - |
Name: Inner capsid protein VP2
Number of copies: 10
UniProtKB UniProtKB PDBe-KB
Molecular function:
RNA binding
Cellular location:
viral inner capsid viral nucleocapsid T=2 icosahedral viral capsid
Number of copies: 10
UniProtKB UniProtKB PDBe-KB
Domains
Gene Ontology [4]
Molecular function:
RNA binding
Cellular location:
viral inner capsid viral nucleocapsid T=2 icosahedral viral capsid
Natural source
Organism: Rotavirus A
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 103 kDa | - |
Name: URIDINE 5'-TRIPHOSPHATE
HET code: UTP
Number of copies: 1
ChEMBL ChEBI DrugBank
HET code: UTP
Number of copies: 1
ChEMBL ChEBI DrugBank
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 484 Da | - |
Name: GUANOSINE-5'-TRIPHOSPHATE
HET code: GTP
Number of copies: 1
ChEMBL ChEBI DrugBank
HET code: GTP
Number of copies: 1
ChEMBL ChEBI DrugBank
Molecular weight
| Experimental | Theoretical | Method |
|---|---|---|
| - | 523 Da | - |
