EMD-12585
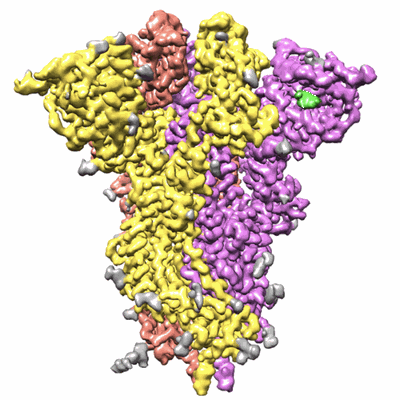
Trimeric SARS-CoV-2 spike ectodomain in complex with biliverdin (closed conformation)
EMD-12585
Single-particle3.36 Å
 Deposition: 09/03/2021
Deposition: 09/03/2021Map released: 28/04/2021
Last modified: 09/06/2021
Concentration: 0.6
mg/mL
Details: 0.6 mg/ml final concentration in TBSE supplemented with 0.1% n-octylglucoside) with 25 micro M biliverdin
Details: 0.6 mg/ml final concentration in TBSE supplemented with 0.1% n-octylglucoside) with 25 micro M biliverdin
Buffer
pH: 8.0
Details: 150 mM NaCl, 1 mM ethylenediaminetetraacetic acid (EDTA), 25 mM Tris-HCl, pH 8.0
Details: 150 mM NaCl, 1 mM ethylenediaminetetraacetic acid (EDTA), 25 mM Tris-HCl, pH 8.0
Grid
Vitrification
Cryogen name: ETHANE
Microscope: FEI TITAN KRIOS
Illumination mode: FLOOD BEAM
Imaging mode: BRIGHT FIELD
Electron source: FIELD EMISSION GUN
Acceleration voltage: 300 kV
Illumination mode: FLOOD BEAM
Imaging mode: BRIGHT FIELD
Electron source: FIELD EMISSION GUN
Acceleration voltage: 300 kV
Image Recording
[1]
Detector model:
FEI FALCON III (4k x 4k)
Detector mode: COUNTING
Number of grids: 1
Number of real images: 15962
Average electron dose per image: 33.0 e/Å2
Details: A total of 15962 movies were recorded with a calibrated pixel size of 1.09 Angstrom and a total electron exposure of 33 electrons per square Angstrom, spread over 30 frames in single electron counting mode.
Detector mode: COUNTING
Number of grids: 1
Number of real images: 15962
Average electron dose per image: 33.0 e/Å2
Details: A total of 15962 movies were recorded with a calibrated pixel size of 1.09 Angstrom and a total electron exposure of 33 electrons per square Angstrom, spread over 30 frames in single electron counting mode.
Final
reconstruction
Resolution: 3.36
Å
(
BY AUTHOR)
Resolution method: FSC 0.143 CUT-OFF
Number of images used: 78073
Resolution method: FSC 0.143 CUT-OFF
Number of images used: 78073
⌯ Applied Symmetry
Point group:
C3
Software
[1]
| Name | Version | Details |
|---|---|---|
| CryoSPARC | 2 | - |
⦨ Initial angle
assignment
Type:
ANGULAR RECONSTITUTION
⦩ Final angle assignment
Type:
ANGULAR RECONSTITUTION
Format: CCP4
Data type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
Annotation details: Full map, cryosparc local filter
Data type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
Annotation details: Full map, cryosparc local filter
⬡ Geometry
| X | Y | Z | |
|---|---|---|---|
| Dimensions | 320 | 320 | 320 |
| Origin | 0 | 0 | 0 |
| Spacing | 320 | 320 | 320 |
| Voxel size | 1.09 Å | 1.09 Å | 1.09 Å |
Contour list
| Primary | Level | Source |
|---|---|---|
| True | 0.72 | AUTHOR |
