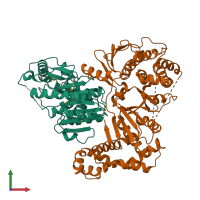
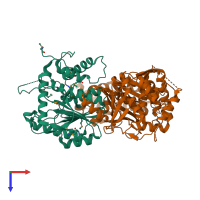
Function and Biology Details
Reactions catalysed:
S-adenosyl-L-methionine + a 5'-(N(7)-methyl 5'-triphosphoguanosine)-(ribonucleotide)-[mRNA] = S-adenosyl-L-homocysteine + a 5'-(N(7)-methyl 5'-triphosphoguanosine)-(2'-O-methyl-ribonucleotide)-[mRNA]
ATP + RNA(n) = diphosphate + RNA(n+1)
Biochemical function:
Biological process:
Cellular component:
- not assigned
Sequence domains:
- Poly(A) polymerase catalytic subunit, Poxvirus
- Poxvirus cap-specific nucleoside 2-O-methyltransferase
- Poxvirus poly(A) polymerase, catalytic subunit, C-terminal
- Poly(A) polymerase, nucleotidyltransferase domain superfamily, Poxvirus
- Poly(A) polymerase, nucleotidyltransferase domain, Poxvirus
- Poxvirus poly(A) polymerase, catalytic subunit, N-terminal
- Poly(A) polymerase catalytic subunit superfamily
- Poxvirus poly(A) polymerase, catalytic subunit, N-terminal domain superfamily
3 more domains
Structure domains:
Structure analysis Details
Assembly composition:
hetero dimer (preferred)
Assembly name:
PDBe Complex ID:
PDB-CPX-149942 (preferred)
Entry contents:
2 distinct polypeptide molecules
Macromolecules (2 distinct):