 |
PDBsum entry 2osx

|

|

|
|
 |
Contents |
 |
|
|

|

|
|
|
|
|
|
|
|
|
|
* Residue conservation analysis
|
|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class:
|
 |
E.C.3.2.1.123
- endoglycosylceramidase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
an oligoglycosyl-(1->4)-beta-D-glucosyl-(1<->1)-ceramide + H2O = an oligoglycosyl-(1->4)-D-glucose + an N-acyl-sphingoid base
|
 |
 |
 |
 |
 |
Oligoglycosylglucosyl-(1<->1)-ceramide
|
+
|
H(2)O
|
=
|
ceramide
Bound ligand (Het Group name = )
matches with 40.62% similarity
|
+
|
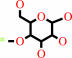 oligoglycosylglucose
oligoglycosylglucose
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
DOI no:
|
J Biol Chem
282:14300-14308
(2007)
|
|
PubMed id:
|
|
|
|

|
| |
|
Structural and mechanistic analyses of endo-glycoceramidase II, a membrane-associated family 5 glycosidase in the Apo and GM3 ganglioside-bound forms.
|
|
M.E.Caines,
M.D.Vaughan,
C.A.Tarling,
S.M.Hancock,
R.A.Warren,
S.G.Withers,
N.C.Strynadka.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
endo-Glycoceramidase, a membrane-associated family 5 glycosidase, deviates from
the typical polysaccharide substrate specificity of other soluble members of the
family, preferentially hydrolyzing glycosidic linkages between the
oligosaccharide and ceramide moieties of gangliosides. Here we report the first
x-ray crystal structures of an endo-glycoceramidase from Rhodococcus sp., in the
apo form, in complex with the ganglioside G(M3) (Svennerholm ganglioside
nomenclature (Svennerholm, L. (1964) J. Lipid Res. 5, 145-155)), and trapped as
a glycosyl-enzyme intermediate. These snapshots provide the first molecular
insight into enzyme recognition and association with gangliosides, revealing the
structural adaptations necessary for glycosidase-catalyzed hydrolysis and
detailing a novel ganglioside binding topology. Consistent with the chemical
duality of the substrate, the active site of endo-glycoceramidase is split into
a wide, polar cavity to bind the polyhydroxylated oligosaccharide moiety and a
narrow, hydrophobic tunnel to bind the ceramide lipid chains. The specific
interactions with the ceramide polar head group manifest a surprising aglycone
specificity, an observation substantiated by our kinetic analyses. Collectively,
the reported structural and kinetic data provide insight toward rational
redesign of the synthetic glycosynthase mutant of endo-glycoceramidase to enable
facile synthesis of nonnatural, therapeutically useful gangliosides.

|

|
|
|

|
| |
Selected figure(s)
|
|

|
| |
 |
 |

|
 |

|
 |
Figure 2.
FIGURE 2. a, the structure of the EGC monomer. b, the
electrostatic surface potential of EGC (red, electronegative;
blue, electropositive; contoured from -15 to 1 kT/e). c, the
hydrophobic surface potential of EGC (green, hydrophobic; white,
polar). d, the structure of the  -(1,4)-glucanase from
Bacillus agaradherans, Cel5A (Protein Data Bank code 2A3H). e,
the electrostatic surface potential of Cel5A (red,
electronegative; blue, electropositive; contoured from -20 to 1
kT/e). Bound ligands, G[M3] (a and b) and cellobiose (d and e),
are shown as ball-and-stick representations in yellow. -(1,4)-glucanase from
Bacillus agaradherans, Cel5A (Protein Data Bank code 2A3H). e,
the electrostatic surface potential of Cel5A (red,
electronegative; blue, electropositive; contoured from -20 to 1
kT/e). Bound ligands, G[M3] (a and b) and cellobiose (d and e),
are shown as ball-and-stick representations in yellow.
|
 |
Figure 3.
FIGURE 3. a, electron density for the bound G[M3].AnmF[o] -
DF[c] (36) electron density map, calculated after random model
perturbation and refinement with G[M3] atoms omitted, is shown
contoured around the G[M3] at 2.5  in red. b, a surface
representation depicting the G[M3] binding site. G[M3] is shown
as a ball-and-stick representation in yellow, surrounded by its
ligands in the active site of EGC. c, a schematic representation
of polar, close contacts involved in the binding of G[M3]. Water
molecules are represented by gray spheres. in red. b, a surface
representation depicting the G[M3] binding site. G[M3] is shown
as a ball-and-stick representation in yellow, surrounded by its
ligands in the active site of EGC. c, a schematic representation
of polar, close contacts involved in the binding of G[M3]. Water
molecules are represented by gray spheres.
|
 |
|
|

|
| |
The above figures are
reprinted
by permission from the ASBMB:
J Biol Chem
(2007,
282,
14300-14308)
copyright 2007.
|
|
| |
Figures were
selected
by an automated process.
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Literature references that cite this PDB file's key reference
|
|
 |
| |
PubMed id
|
 |
Reference
|
 |
|
|
|
 |
H.Karlsson,
A.Halim,
and
S.Teneberg
(2010).
Differentiation of glycosphingolipid-derived glycan structural isomers by liquid chromatography/mass spectrometry.
|
| |
Glycobiology,
20,
1103-1116.
|
 |
|
|
|
|
 |
L.X.Wang,
and
W.Huang
(2009).
Enzymatic transglycosylation for glycoconjugate synthesis.
|
| |
Curr Opin Chem Biol,
13,
592-600.
|
 |
|
|
|
|
 |
O.Kanie,
A.Kurimoto,
Y.Kanie,
S.Daikoku,
A.Ohtake,
and
K.Suzuki
(2009).
Analysis of behavior of sodiated sugar hemiacetals under low-energy collision-induced dissociation conditions and application to investigating mutarotation and mechanism of a glycosidase.
|
| |
Proc Jpn Acad Ser B Phys Biol Sci,
85,
204-215.
|
 |
|
|
|
|
 |
F.A.Shaikh,
and
S.G.Withers
(2008).
Teaching old enzymes new tricks: engineering and evolution of glycosidases and glycosyl transferases for improved glycoside synthesis.
|
| |
Biochem Cell Biol,
86,
169-177.
|
 |
|
|
|
|
 |
Y.Kacher,
B.Brumshtein,
S.Boldin-Adamsky,
L.Toker,
A.Shainskaya,
I.Silman,
J.L.Sussman,
and
A.H.Futerman
(2008).
Acid beta-glucosidase: insights from structural analysis and relevance to Gaucher disease therapy.
|
| |
Biol Chem,
389,
1361-1369.
|
 |
|
PDB code:
|
 |
|
|
 |
 |
|
The most recent references are shown first.
Citation data come partly from CiteXplore and partly
from an automated harvesting procedure. Note that this is likely to be
only a partial list as not all journals are covered by
either method. However, we are continually building up the citation data
so more and more references will be included with time.
Where a reference describes a PDB structure, the PDB
code is
shown on the right.
|
 |

