 |
PDBsum entry 2kaf

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Viral protein, RNA binding protein
|
PDB id
|
|

|

|
2kaf
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
|

|
|
PDB id:
|
 |
|
 |
| Name: |
 |
Viral protein, RNA binding protein
|
 |
|
Title:
|
 |
Solution structure of the sars-unique domain-c from the nonstructural protein 3 (nsp3) of the severe acute respiratory syndrome coronavirus
|
|
Structure:
|
 |
Non-structural protein 3. Chain: a. Fragment: sars-unique domain-c, residues 1473-1532. Synonym: replicase polyprotein 1a, pp1a, orf1a polyprotein, nsp3, papain-like proteinase, pl-pro, pl2-pro. Engineered: yes
|
|
Source:
|
 |
Sars coronavirus. Sars-cov. Organism_taxid: 227859. Gene: 1a. Expressed in: escherichia coli. Expression_system_taxid: 562.
|
|
NMR struc:
|
 |
20 models
|
 |
|
Authors:
|
 |
M.A.Johnson,B.Mohanty,B.Pedrini,P.Serrano,A.Chatterjee,T.Herrmann, J.Joseph,K.Saikatendu,I.A.Wilson,M.J.Buchmeier,P.Kuhn,K.Wuthrich, Joint Center For Structural Genomics (Jcsg)
|
|
Key ref:
|
 |
M.A.Johnson
et al.
(2010).
SARS coronavirus unique domain: three-domain molecular architecture in solution and RNA binding.
J Mol Biol,
400,
724-742.
PubMed id:
|
 |
|
Date:
|
 |
|
05-Nov-08
|
Release date:
|
25-Nov-08
|
|
|

|

|
|
PROCHECK
|

|

|
|
|
Headers
|
 |
|
|
References
|
|
|
|

|

|
|
P0C6U8
(R1A_CVHSA) -
Replicase polyprotein 1a from Severe acute respiratory syndrome coronavirus
|
|

|

|
Seq:
Struc:
|
 |
 |
 |
4382 a.a.
67 a.a.*
|

|

|

|

|
|

|

|

|
 |
 |

|
|
Key: |
 |
PfamA domain |
 |
 |
 |
Secondary structure |
 |
 |
CATH domain |
 |
|
*
PDB and UniProt seqs differ
at 1 residue position (black
cross)
|
|

|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class 1:
|
 |
E.C.2.7.7.50
- mRNA guanylyltransferase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
a 5'-end diphospho-ribonucleoside in mRNA + GTP + H+ = a 5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA + diphosphate
|
 |
 |
 |
 |
 |
5'-end diphospho-ribonucleoside in mRNA
|
+
|
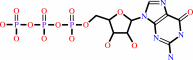 GTP
GTP
|
+
|
H(+)
|
=
|
5'-end (5'-triphosphoguanosine)-ribonucleoside in mRNA
|
+
|
 diphosphate
diphosphate
|
|
 |
 |
 |
 |
 |
 |
 |
 |
Enzyme class 2:
|
 |
E.C.3.4.19.12
- ubiquitinyl hydrolase 1.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
Thiol-dependent hydrolysis of ester, thiolester, amide, peptide and isopeptide bonds formed by the C-terminal Gly of ubiquitin (a 76-residue protein attached to proteins as an intracellular targeting signal).
|
 |
 |
 |
 |
 |
Enzyme class 3:
|
 |
E.C.3.4.22.-
- ?????
|
|
 |
 |
 |
 |
 |
Enzyme class 4:
|
 |
E.C.3.4.22.69
- Sars coronavirus main proteinase.
|
|
 |
 |
 |
 |
 |
 |
 |
|
Note, where more than one E.C. class is given (as above), each may
correspond to a different protein domain or, in the case of polyprotein
precursors, to a different mature protein.
|
|
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
|
J Mol Biol
400:724-742
(2010)
|
|
PubMed id:
|
|
|
|

|
| |
|
SARS coronavirus unique domain: three-domain molecular architecture in solution and RNA binding.
|
|
M.A.Johnson,
A.Chatterjee,
B.W.Neuman,
K.Wüthrich.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
The nonstructural protein 3 (nsp3) of the severe acute respiratory syndrome
coronavirus (SARS-CoV) includes a "SARS-unique region" (SUD) consisting of three
globular domains separated by short linker peptide segments. This paper reports
NMR structure determinations of the C-terminal domain (SUD-C) and of a
two-domain construct (SUD-MC) containing the middle domain (SUD-M) and the
C-terminal domain, and NMR data on the conformational states of the N-terminal
domain (SUD-N) and the SUD-NM two-domain construct. Both SUD-N and SUD-NM are
monomeric and globular in solution, and in SUD-NM there is high mobility in the
two-residue interdomain linking sequence, with no preferred relative orientation
of the two domains. SUD-C adopts a frataxin-like fold and has structural
similarity to DNA-binding domains of DNA-modifying enzymes. The structures of
both SUD-M (previously determined) and SUD-C (from the present study) are
maintained in SUD-MC, where the two domains are flexibly linked. Gel shift
experiments showed that both SUD-C and SUD-MC bind to single-stranded RNA and
recognize purine bases more strongly than pyrimidine bases, whereby SUD-MC binds
to a more restricted set of purine-containing RNA sequences than SUD-M. NMR
chemical shift perturbation experiments with observation of the (15)N-labeled
proteins further resulted in the delineation of the RNA binding sites, i.e., in
SUD-M a positively charged surface area with a pronounced cavity, and in SUD-C
several residues of an antiparallel beta-sheet. Overall, the present data
provide evidence for molecular mechanisms involving concerted actions of SUD-M
and SUD-C, which result in specific RNA-binding that might be unique to the SUD,
and thus to the SARS-CoV.

|

|
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

