 |
PDBsum entry 2j4f

|

|

|
|
 |
Contents |
 |
|
|

|

|
|
|
|
|
|
|
|
* Residue conservation analysis
|
|
|
|
 |
|
|
 |
 |
 |
 |
Enzyme class:
|
 |
E.C.3.1.1.7
- acetylcholinesterase.
|
|
 |
 |
 |
 |
 |
Reaction:
|
 |
acetylcholine + H2O = choline + acetate + H+
|
 |
 |
 |
 |
 |
 acetylcholine
acetylcholine
|
+
|
H2O
|
=
|
 choline
choline
|
+
|
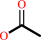 acetate
acetate
|
+
|
H(+)
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Molecule diagrams generated from .mol files obtained from the
KEGG ftp site
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
|
| |
|
DOI no:
|
Biochemistry
33:14407-14418
(1994)
|
|
PubMed id:
|
|
|
|

|
| |
|
A metastable state of Torpedo californica acetylcholinesterase generated by modification with organomercurials.
|
|
D.I.Kreimer,
E.A.Dolginova,
M.Raves,
J.L.Sussman,
I.Silman,
L.Weiner.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
Chemical modification of Torpedo californica acetylcholinesterase by various
sulfhydryl reagents results in its conversion to one of two principal states.
One of these states, viz., that produced by disulfides and alkylating agents, is
stable. The second state, produced by mercury derivatives, is metastable. At
room temperature, it converts spontaneously, with a half-life of ca. 1 h, to a
stable state similar to that produced by the disulfides and alkylating agents.
Demodification of acetylcholinesterase freshly modified by mercurials, by its
exposure to reduced glutathione, causes rapid release of the bound mercurial,
with concomitant recovery of most of the enzymic activity of the native enzyme.
In contrast, similar demodification of acetylcholinesterase modified by
disulfides yields no detectable recovery of enzymic activity. Spectroscopic
measurements, employing CD, intrinsic fluorescence, and binding of
1-anilino-8-naphthalenesulfonate, show that the state produced initially by
mercurials is "native-like", whereas that produced by disulfides and alkylating
agents, and after prolonged incubation of the mercurial-modified enzyme, is
partially unfolded and displays many of the features of the "molten globule"
state. Arrhenius plots show that the quasi-native state produced by
organomercurials is separated by a low (5 kcal/mol) energy barrier from the
native state, whereas the partially unfolded state is separated from the
quasi-native state by a high energy barrier (ca. 50 kcal/mol). Comparison of the
3D structures of native acetylcholinesterase and of a heavy-atom derivative
obtained with HgAc2 suggests that the mercurial-modified enzyme may be
stabilized by additional interactions of the mercury atom attached to the free
thiol group of Cys231, specifically with Ser228O gamma with the main-chain
nitrogen and carbonyl oxygen of the same serine residue.

|

|
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
|
Literature references that cite this PDB file's key reference
|
|
 |
| |
PubMed id
|
 |
Reference
|
 |
|
|
|
 |
L.Weiner,
E.Roth,
and
I.Silman
(2011).
Targeted Oxidation of Torpedo californica Acetylcholinesterase by Singlet Oxygen.
|
| |
Photochem Photobiol,
87,
308-316.
|
 |
|
|
|
|
 |
A.Badiou,
J.L.Brunet,
and
L.P.Belzunces
(2007).
Existence of two membrane-bound acetylcholinesterases in the honey bee head.
|
| |
Arch Insect Biochem Physiol,
66,
122-134.
|
 |
|
|
|
|
 |
M.F.Frasco,
J.P.Colletier,
M.Weik,
F.Carvalho,
L.Guilhermino,
J.Stojan,
and
D.Fournier
(2007).
Mechanisms of cholinesterase inhibition by inorganic mercury.
|
| |
FEBS J,
274,
1849-1861.
|
 |
|
PDB code:
|
 |
|
|
|
|
|
 |
C.B.Millard,
V.L.Shnyrov,
S.Newstead,
I.Shin,
E.Roth,
I.Silman,
and
L.Weiner
(2003).
Stabilization of a metastable state of Torpedo californica acetylcholinesterase by chemical chaperones.
|
| |
Protein Sci,
12,
2337-2347.
|
 |
|
|
|
|
 |
I.Shin,
E.Wachtel,
E.Roth,
C.Bon,
I.Silman,
and
L.Weiner
(2002).
Thermal denaturation of Bungarus fasciatus acetylcholinesterase: Is aggregation a driving force in protein unfolding?
|
| |
Protein Sci,
11,
2022-2032.
|
 |
|
|
|
|
 |
D.I.Kreimer,
K.P.Chai,
and
G.Ferro-Luzzi Ames
(2000).
Nonequivalence of the nucleotide-binding subunits of an ABC transporter, the histidine permease, and conformational changes in the membrane complex.
|
| |
Biochemistry,
39,
14183-14195.
|
 |
|
|
|
|
 |
I.Shin,
D.Kreimer,
I.Silman,
and
L.Weiner
(1997).
Membrane-promoted unfolding of acetylcholinesterase: a possible mechanism for insertion into the lipid bilayer.
|
| |
Proc Natl Acad Sci U S A,
94,
2848-2852.
|
 |
|
|
|
|
 |
M.Peretz,
L.M.Weiner,
and
Y.Burstein
(1997).
Cysteine reactivity in Thermoanaerobacter brockii alcohol dehydrogenase.
|
| |
Protein Sci,
6,
1074-1083.
|
 |
|
|
|
|
 |
I.Shin,
I.Silman,
and
L.M.Weiner
(1996).
Interaction of partially unfolded forms of Torpedo acetylcholinesterase with liposomes.
|
| |
Protein Sci,
5,
42-51.
|
 |
|
 |
 |
|
The most recent references are shown first.
Citation data come partly from CiteXplore and partly
from an automated harvesting procedure. Note that this is likely to be
only a partial list as not all journals are covered by
either method. However, we are continually building up the citation data
so more and more references will be included with time.
Where a reference describes a PDB structure, the PDB
code is
shown on the right.
|
 |

