 |
PDBsum entry 1k0y

|
|
|

|
 |

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
 |

|

|

|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|

|

|

|

|

|

|
|
Oxygen storage/transport
|
PDB id
|
|

|

|
1k0y
|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|

|
|
 |
Contents |
 |
|
|

|

|
|
|
|
|
|
|
|
* Residue conservation analysis
|
|
|

|
|
PDB id:
|
 |
|
 |
| Name: |
 |
Oxygen storage/transport
|
 |
|
Title:
|
 |
X-ray crystallographic analyses of symmetrical allosteric effectors of hemoglobin. Compounds designed to link primary and secondary binding sites
|
|
Structure:
|
 |
Hemoglobin alpha chain. Chain: a, c. Hemoglobin beta chain. Chain: b, d
|
|
Source:
|
 |
Homo sapiens. Human. Organism_taxid: 9606. Tissue: blood. Tissue: blood
|
|
Biol. unit:
|
 |
Tetramer (from
)
|
|
Resolution:
|
 |
|
1.87Å
|
R-factor:
|
0.163
|
R-free:
|
0.189
|
|
|
Authors:
|
 |
M.K.Safo,T.Boyiri,J.C.Burnett,R.Danso-Danquah,C.M.Moure,G.S.Joshi, D.J.Abraham
|
Key ref:

|
 |
M.K.Safo
et al.
(2002).
X-ray crystallographic analyses of symmetrical allosteric effectors of hemoglobin: compounds designed to link primary and secondary binding sites.
Acta Crystallogr D Biol Crystallogr,
58,
634-644.
PubMed id:
DOI:

|
 |
|
Date:
|
 |
|
21-Sep-01
|
Release date:
|
03-Oct-01
|
|
|

|

|
|
PROCHECK
|

|

|
|
|
Headers
|
 |
|
|
References
|
|
|
|
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |

|

|

|
| |
|
DOI no:
|
Acta Crystallogr D Biol Crystallogr
58:634-644
(2002)
|
|
PubMed id:
|
|
|
|

|
| |
|
X-ray crystallographic analyses of symmetrical allosteric effectors of hemoglobin: compounds designed to link primary and secondary binding sites.
|
|
M.K.Safo,
T.Boyiri,
J.C.Burnett,
R.Danso-Danquah,
C.M.Moure,
G.S.Joshi,
D.J.Abraham.
|
|
|

|
| |
ABSTRACT
|
|

|
| |

|
The rational design and X-ray crystallographic analyses of two symmetrical
allosteric effectors of hemoglobin (Hb) are reported. Compound design was
directed by the previously solved co-crystal structure of one of the most potent
allosteric effectors of Hb,
2-[4-[(3,5-dichlorophenylcarbamoyl)-methyl]-phenoxy]-2-methylpropionic acid
(RSR4), which revealed two distinct binding sites for this compound in the Hb
central water cavity. The primary binding site has been observed for all
compounds of this structural class, which stabilize deoxy Hb by engaging in
inter-dimer contacts with three of the four protein subunits. Interactions at
the secondary binding site of RSR4 occur primarily between the beta(1) and
beta(2) subunits and serve to further constrain the deoxy state. Based on these
observations, it was hypothesized that compounds with the ability to
simultaneously span and link both of these sites would possess increased
potency, but at a lower molar concentration than RSR4. Two symmetrical compounds
were designed and synthesized based on this hypothesis. The symmetrical effector
approach was taken to minimize the number of compound orientations needed to
successfully bind at either of the distinct allosteric sites. X-ray
crystallographic analyses of these two effectors in complex with Hb revealed
that they successfully spanned the RSR4 primary and secondary binding sites.
However, the designed compounds interacted with the secondary binding site in
such a way that intra-dimer, as opposed to inter-dimer, interactions were
generated. In agreement with these observations, in vitro evaluation of the
symmetrical effectors in Hb solution indicated that neither compound possessed
the potency of RSR4. A detailed analysis of symmetrical effector-Hb contacts and
comparisons with the binding contacts of RSR4 are discussed.

|

|
|
|

|
| |
Selected figure(s)
|
|

|
| |
 |
 |

|
 |

|
 |
Figure 5.
Figure 5 Stereoviews of allosteric sites of effectors (a) RSR4,
(b) TB5-27 and (c) TB5-39. Protein and effector atoms are shown
as sticks and structural waters are red spheres. The Hb  [1]
subunit is light blue, the [1]
subunit is light blue, the  [2]
subunit is magenta, the [2]
subunit is magenta, the 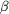 [1]
subunit is orange and the [1]
subunit is orange and the 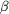 [2
]subunit is red. C atoms of allosteric effectors are yellow, O
atoms are red and N atoms are blue. Dashed black lines indicate
hydrogen bonds. For clarity, not all Hb residues lining the
allosteric binding sites are shown. Note, for visualization
purposes, and owing to secondary-site binding orientation
differences between RSR4 and the symmetrical effectors, Hb
residue labels and interaction distances for RSR4 in Fig. 5-(a)
are labeled on the opposite side of the dimer-dimer interface
with respect to labeled residues and interaction distances in
the TB5-27-Hb complex (Fig. 5-b) and the TB5-39-Hb complex (Fig.
5-c). For clarity of description, textual reference to RSR4-Hb
residue contacts follow dimer-dimer interface residue labeling
as pictured for TB5-27 and TB5-39. Importantly, the described
RSR4 interactions in the text correspond to equivalent
interactions made by this effector at its symmetry-related
binding sites and are the mirror image of the contacts labeled
in Fig. 5-(a).-. [2
]subunit is red. C atoms of allosteric effectors are yellow, O
atoms are red and N atoms are blue. Dashed black lines indicate
hydrogen bonds. For clarity, not all Hb residues lining the
allosteric binding sites are shown. Note, for visualization
purposes, and owing to secondary-site binding orientation
differences between RSR4 and the symmetrical effectors, Hb
residue labels and interaction distances for RSR4 in Fig. 5-(a)
are labeled on the opposite side of the dimer-dimer interface
with respect to labeled residues and interaction distances in
the TB5-27-Hb complex (Fig. 5-b) and the TB5-39-Hb complex (Fig.
5-c). For clarity of description, textual reference to RSR4-Hb
residue contacts follow dimer-dimer interface residue labeling
as pictured for TB5-27 and TB5-39. Importantly, the described
RSR4 interactions in the text correspond to equivalent
interactions made by this effector at its symmetry-related
binding sites and are the mirror image of the contacts labeled
in Fig. 5-(a).-.
|
 |
Figure 6.
Figure 6 Stereoviews of symmetrically related pairs of
allosteric effectors bound in the Hb central water cavity.
Protein residues are shown as light blue sticks. RSR4 is blue,
RSR13 is orange, TB5-27 is yellow and TB5-39 is red.
Secondary-site binding for RSR4 is inter-dimer, while an
intra-dimer binding mode is adopted by the TB5 compounds.
|
 |
|
|

|
| |
The above figures are
reprinted
by permission from the IUCr:
Acta Crystallogr D Biol Crystallogr
(2002,
58,
634-644)
copyright 2002.
|
|
| |
Figures were
selected
by an automated process.
|
|

|

|
|
 |
 |
|
 |
 |
 |
 |
 |
 |
 |
 |
 |
 |
|

