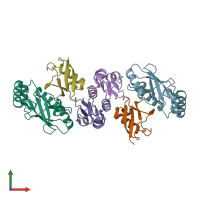
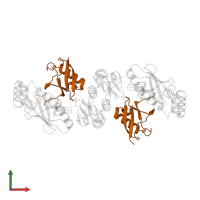
Assemblies
Multimeric state:
hetero hexamer
Accessible surface area:
26387.58 Å2
Buried surface area:
9005.69 Å2
Dissociation area:
4,495.24
Å2
Dissociation energy (ΔGdiss):
-8.36
kcal/mol
Dissociation entropy (TΔSdiss):
57.76
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-143298
Macromolecules
Chains: A, B
Length: 147 amino acids
Theoretical weight: 16.85 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam: Ubiquitin-conjugating enzyme
InterPro:
Length: 147 amino acids
Theoretical weight: 16.85 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 P62837 (Residues: 1-147; Coverage: 100%)
P62837 (Residues: 1-147; Coverage: 100%)
Pfam: Ubiquitin-conjugating enzyme
InterPro:
Chains: C, D
Length: 79 amino acids
Theoretical weight: 8.86 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Ubiquitin family
InterPro:
Length: 79 amino acids
Theoretical weight: 8.86 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P0CG47 (Residues: 153-228; Coverage: 33%)
P0CG47 (Residues: 153-228; Coverage: 33%)
Pfam: Ubiquitin family
InterPro:
Chains: F, G
Length: 123 amino acids
Theoretical weight: 13.11 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam: RING-type zinc-finger
InterPro:
Length: 123 amino acids
Theoretical weight: 13.11 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 P36406 (Residues: 1-123; Coverage: 21%)
P36406 (Residues: 1-123; Coverage: 21%)
Pfam: RING-type zinc-finger
InterPro: