Assemblies
Assembly Name:
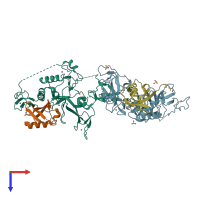
E3 ubiquitin-protein ligase parkin and Ubiquitin
Multimeric state:
hetero dimer
Accessible surface area:
17937.7 Å2
Buried surface area:
2714.87 Å2
Dissociation area:
1,171.56
Å2
Dissociation energy (ΔGdiss):
10.43
kcal/mol
Dissociation entropy (TΔSdiss):
11.24
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-122708
Assembly Name:
E3 ubiquitin-protein ligase parkin and Ubiquitin
Multimeric state:
hetero dimer
Accessible surface area:
19379.93 Å2
Buried surface area:
3234.69 Å2
Dissociation area:
1,196.58
Å2
Dissociation energy (ΔGdiss):
9.51
kcal/mol
Dissociation entropy (TΔSdiss):
11.27
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-122708
Macromolecules
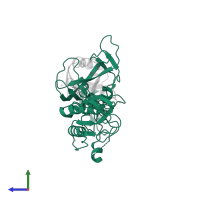
Chains: A, C
Length: 321 amino acids
Theoretical weight: 36.1 KDa
Source organism: Pediculus humanus corporis
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 321 amino acids
Theoretical weight: 36.1 KDa
Source organism: Pediculus humanus corporis
Expression system: Escherichia coli
UniProt:
- Canonical:
 E0VIU9 (Residues: 141-461; Coverage: 70%)
E0VIU9 (Residues: 141-461; Coverage: 70%)
Pfam:
InterPro:
- Parkin, RING/Ubox like zinc-binding domain
- E3 ubiquitin-protein ligase parkin
- E3 ubiquitin ligase RBR family
- RING/Ubox-like zinc-binding domain
- TRIAD supradomain
- E3 ubiquitin-protein ligase parkin, RING finger, HC subclass
- E3 ubiquitin-protein ligase parkin, BRcat domain
- IBR domain
- E3 ubiquitin-protein ligase parkin, Rcat domain
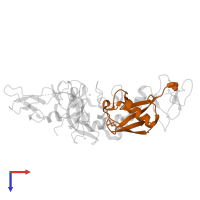
Chains: B, D
Length: 76 amino acids
Theoretical weight: 8.64 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Ubiquitin family
InterPro:
Length: 76 amino acids
Theoretical weight: 8.64 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P0CG47 (Residues: 153-228; Coverage: 33%)
P0CG47 (Residues: 153-228; Coverage: 33%)
Pfam: Ubiquitin family
InterPro: