Assemblies
Assembly Name:
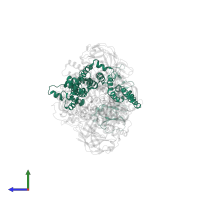
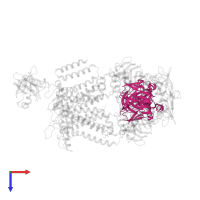
Enzyme IIA-maltose transport inhibitory complex
Multimeric state:
hetero hexamer
Accessible surface area:
74998.7 Å2
Buried surface area:
22127.85 Å2
Dissociation area:
3,062.88
Å2
Dissociation energy (ΔGdiss):
-0.79
kcal/mol
Dissociation entropy (TΔSdiss):
26.01
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-136376
Assembly Name:
Enzyme IIA-maltose transport inhibitory complex
Multimeric state:
hetero hexamer
Accessible surface area:
75258.43 Å2
Buried surface area:
22140.43 Å2
Dissociation area:
5,466.42
Å2
Dissociation energy (ΔGdiss):
16.03
kcal/mol
Dissociation entropy (TΔSdiss):
41.19
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-136376
Macromolecules
Chains: F, H
Length: 514 amino acids
Theoretical weight: 57.05 KDa
Source organism: Escherichia coli K-12
Expression system: Escherichia coli
UniProt:
Pfam:
Length: 514 amino acids
Theoretical weight: 57.05 KDa
Source organism: Escherichia coli K-12
Expression system: Escherichia coli
UniProt:
- Canonical:
 P02916 (Residues: 1-514; Coverage: 100%)
P02916 (Residues: 1-514; Coverage: 100%)
Pfam:
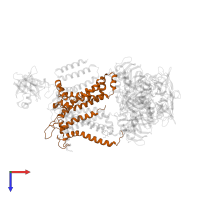
- MalF, N-terminal region, transmembrane helices
- Maltose transport system permease protein MalF P2 domain
- Binding-protein-dependent transport system inner membrane component
- MalF, N-terminal
- Maltose transport system permease protein MalF, P2 domain
- Maltose transport system permease protein MalF, P2 domain superfamily
- MetI-like superfamily
- ABC transporter type 1, transmembrane domain MetI-like
Chains: G, I
Length: 296 amino acids
Theoretical weight: 32.25 KDa
Source organism: Escherichia coli K-12
Expression system: Escherichia coli
UniProt:
Pfam: Binding-protein-dependent transport system inner membrane component
InterPro:
CATH: MetI-like
Length: 296 amino acids
Theoretical weight: 32.25 KDa
Source organism: Escherichia coli K-12
Expression system: Escherichia coli
UniProt:
- Canonical:
 P68183 (Residues: 1-296; Coverage: 100%)
P68183 (Residues: 1-296; Coverage: 100%)
Pfam: Binding-protein-dependent transport system inner membrane component
InterPro:
CATH: MetI-like
Chains: A, B, C, D
Length: 381 amino acids
Theoretical weight: 42.18 KDa
Source organism: Escherichia coli K-12
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 381 amino acids
Theoretical weight: 42.18 KDa
Source organism: Escherichia coli K-12
Expression system: Escherichia coli
UniProt:
- Canonical:
 P68187 (Residues: 1-371; Coverage: 100%)
P68187 (Residues: 1-371; Coverage: 100%)
Pfam:
InterPro:
- P-loop containing nucleoside triphosphate hydrolase
- ABC transporter, ATP-binding protein MalK/UgpC-like
- ABC transporter-like, ATP-binding domain
- ABC transporter, maltose/maltodextrin import, MalK-like
- AAA+ ATPase domain
- ABC transporter-like, conserved site
- MalK-like, OB fold domain
- Molybdate/tungstate binding, C-terminal
- Nucleic acid-binding, OB-fold
Chains: M, N, O, P
Length: 172 amino acids
Theoretical weight: 18.55 KDa
Source organism: Escherichia coli K-12
Expression system: Escherichia coli
UniProt:
Pfam: phosphoenolpyruvate-dependent sugar phosphotransferase system, EIIA 1
InterPro:
CATH: Glucose Permease (Domain IIA)
Length: 172 amino acids
Theoretical weight: 18.55 KDa
Source organism: Escherichia coli K-12
Expression system: Escherichia coli
UniProt:
- Canonical:
 P69783 (Residues: 1-169; Coverage: 100%)
P69783 (Residues: 1-169; Coverage: 100%)
Pfam: phosphoenolpyruvate-dependent sugar phosphotransferase system, EIIA 1
InterPro:
CATH: Glucose Permease (Domain IIA)



![The deposited structure of PDB entry <span class='highlight'>4jbw</span> coloured by chain, front view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>2 copies of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalF</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalG</span>;</li> <li class='image_legend_li'>4 copies of <span class='highlight'>Maltose/maltodextrin import ATP-binding protein MalK</span>;</li> <li class='image_legend_li'>4 copies of <span class='highlight'>PTS system glucose-specific EIIA component</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>(1R)-2-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL (11E)-OCTADEC-11-ENOATE</span>.</li></ul> PDB entry 4jbw coloured by chain, front view.](https://www.ebi.ac.uk/pdbe/static/entry/4jbw_deposited_chain_front_image-200x200.png)
![The deposited structure of PDB entry <span class='highlight'>4jbw</span> coloured by chain, side view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>2 copies of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalF</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalG</span>;</li> <li class='image_legend_li'>4 copies of <span class='highlight'>Maltose/maltodextrin import ATP-binding protein MalK</span>;</li> <li class='image_legend_li'>4 copies of <span class='highlight'>PTS system glucose-specific EIIA component</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>(1R)-2-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL (11E)-OCTADEC-11-ENOATE</span>.</li></ul> PDB entry 4jbw coloured by chain, side view.](https://www.ebi.ac.uk/pdbe/static/entry/4jbw_deposited_chain_side_image-200x200.png)
![The deposited structure of PDB entry <span class='highlight'>4jbw</span> coloured by chain, top view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>2 copies of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalF</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalG</span>;</li> <li class='image_legend_li'>4 copies of <span class='highlight'>Maltose/maltodextrin import ATP-binding protein MalK</span>;</li> <li class='image_legend_li'>4 copies of <span class='highlight'>PTS system glucose-specific EIIA component</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>(1R)-2-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL (11E)-OCTADEC-11-ENOATE</span>.</li></ul> PDB entry 4jbw coloured by chain, top view.](https://www.ebi.ac.uk/pdbe/static/entry/4jbw_deposited_chain_top_image-200x200.png)
![Hetero hexameric assembly 1 of PDB entry <span class='highlight'>4jbw</span> coloured by chemically distinct molecules, front view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>1 copy of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalF</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalG</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>Maltose/maltodextrin import ATP-binding protein MalK</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>PTS system glucose-specific EIIA component</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>(1R)-2-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL (11E)-OCTADEC-11-ENOATE</span>.</li></ul>} Hetero hexameric assembly 1 of PDB entry 4jbw coloured by chemically distinct molecules, front view.](https://www.ebi.ac.uk/pdbe/static/entry/4jbw_assembly_1_chemically_distinct_molecules_front_image-200x200.png)
![Hetero hexameric assembly 1 of PDB entry <span class='highlight'>4jbw</span> coloured by chemically distinct molecules, side view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>1 copy of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalF</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalG</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>Maltose/maltodextrin import ATP-binding protein MalK</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>PTS system glucose-specific EIIA component</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>(1R)-2-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL (11E)-OCTADEC-11-ENOATE</span>.</li></ul>} Hetero hexameric assembly 1 of PDB entry 4jbw coloured by chemically distinct molecules, side view.](https://www.ebi.ac.uk/pdbe/static/entry/4jbw_assembly_1_chemically_distinct_molecules_side_image-200x200.png)
![Hetero hexameric assembly 1 of PDB entry <span class='highlight'>4jbw</span> coloured by chemically distinct molecules, top view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>1 copy of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalF</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalG</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>Maltose/maltodextrin import ATP-binding protein MalK</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>PTS system glucose-specific EIIA component</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>(1R)-2-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL (11E)-OCTADEC-11-ENOATE</span>.</li></ul>} Hetero hexameric assembly 1 of PDB entry 4jbw coloured by chemically distinct molecules, top view.](https://www.ebi.ac.uk/pdbe/static/entry/4jbw_assembly_1_chemically_distinct_molecules_top_image-200x200.png)
![Hetero hexameric assembly 2 of PDB entry <span class='highlight'>4jbw</span> coloured by chemically distinct molecules, front view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>1 copy of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalF</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalG</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>Maltose/maltodextrin import ATP-binding protein MalK</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>PTS system glucose-specific EIIA component</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>(1R)-2-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL (11E)-OCTADEC-11-ENOATE</span>.</li></ul>} Hetero hexameric assembly 2 of PDB entry 4jbw coloured by chemically distinct molecules, front view.](https://www.ebi.ac.uk/pdbe/static/entry/4jbw_assembly_2_chemically_distinct_molecules_front_image-200x200.png)
![Hetero hexameric assembly 2 of PDB entry <span class='highlight'>4jbw</span> coloured by chemically distinct molecules, side view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>1 copy of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalF</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalG</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>Maltose/maltodextrin import ATP-binding protein MalK</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>PTS system glucose-specific EIIA component</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>(1R)-2-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL (11E)-OCTADEC-11-ENOATE</span>.</li></ul>} Hetero hexameric assembly 2 of PDB entry 4jbw coloured by chemically distinct molecules, side view.](https://www.ebi.ac.uk/pdbe/static/entry/4jbw_assembly_2_chemically_distinct_molecules_side_image-200x200.png)
![Hetero hexameric assembly 2 of PDB entry <span class='highlight'>4jbw</span> coloured by chemically distinct molecules, top view. This structure contains: <ul class='image_legend_ul'><li class='image_legend_li'>1 copy of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalF</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>Maltose/maltodextrin transport system permease protein MalG</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>Maltose/maltodextrin import ATP-binding protein MalK</span>;</li> <li class='image_legend_li'>2 copies of <span class='highlight'>PTS system glucose-specific EIIA component</span>;</li> <li class='image_legend_li'>1 copy of <span class='highlight'>(1R)-2-{[{[(2S)-2,3-DIHYDROXYPROPYL]OXY}(HYDROXY)PHOSPHORYL]OXY}-1-[(PALMITOYLOXY)METHYL]ETHYL (11E)-OCTADEC-11-ENOATE</span>.</li></ul>} Hetero hexameric assembly 2 of PDB entry 4jbw coloured by chemically distinct molecules, top view.](https://www.ebi.ac.uk/pdbe/static/entry/4jbw_assembly_2_chemically_distinct_molecules_top_image-200x200.png)