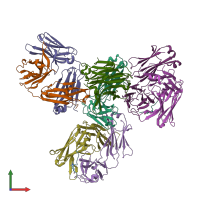
X-ray diffraction
2.53Å resolution
Source organisms:
Primary publication: