Function and Biology Details
Biochemical function:
Biological process:
Cellular component:
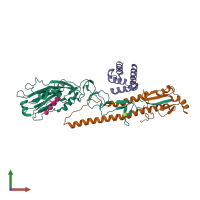
Structure analysis Details
Assembly composition:
hetero nonamer (preferred)
Assembly name:
Hemagglutinin HA1 chain (preferred)
PDBe Complex ID:
PDB-CPX-163961 (preferred)
Entry contents:
3 distinct polypeptide molecules
Macromolecules (4 distinct):
Ligands and Environments
Experiments and Validation Details
X-ray source:
APS BEAMLINE 23-ID-D
Spacegroup:
R32
Expression systems:
- Trichoplusia ni
- Escherichia coli BL21(DE3)