Assemblies
Assembly Name:
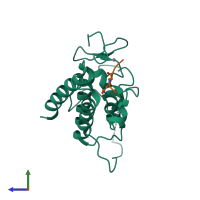
Histone H3.1 and Transcription intermediary factor 1-alpha
Multimeric state:
hetero dimer
Accessible surface area:
10845.7 Å2
Buried surface area:
590.25 Å2
Dissociation area:
295.13
Å2
Dissociation energy (ΔGdiss):
-0.71
kcal/mol
Dissociation entropy (TΔSdiss):
5.73
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-127339
Assembly Name:
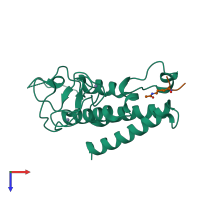
Histone H3.1 and Transcription intermediary factor 1-alpha
Multimeric state:
hetero dimer
Accessible surface area:
10663.77 Å2
Buried surface area:
392.65 Å2
Dissociation area:
196.32
Å2
Dissociation energy (ΔGdiss):
1.3
kcal/mol
Dissociation entropy (TΔSdiss):
2.81
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-127339
Macromolecules
Chains: A, B
Length: 184 amino acids
Theoretical weight: 21.32 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam:
InterPro:
Length: 184 amino acids
Theoretical weight: 21.32 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 O15164 (Residues: 824-1006; Coverage: 17%)
O15164 (Residues: 824-1006; Coverage: 17%)
Pfam:
InterPro:
- Zinc finger, RING/FYVE/PHD-type
- Zinc finger, FYVE/PHD-type
- Zinc finger, PHD-finger
- Zinc finger, PHD-type
- Zinc finger, PHD-type, conserved site
- Bromodomain-like superfamily
- Bromodomain
- Bromodomain, conserved site
Chains: D, E
Length: 9 amino acids
Theoretical weight: 943 Da
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
Length: 9 amino acids
Theoretical weight: 943 Da
Source organism: Homo sapiens
Expression system: Not provided
UniProt:
- Canonical:
 P68431 (Residues: 24-32; Coverage: 7%)
P68431 (Residues: 24-32; Coverage: 7%)