Assemblies
Assembly Name:
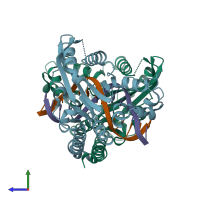
Type II restriction enzyme HincII and DNA
Multimeric state:
hetero tetramer
Accessible surface area:
23351.01 Å2
Buried surface area:
8735.94 Å2
Dissociation area:
2,447.38
Å2
Dissociation energy (ΔGdiss):
45.95
kcal/mol
Dissociation entropy (TΔSdiss):
13.55
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-116203
Macromolecules
Chains: A, B
Length: 317 amino acids
Theoretical weight: 36.59 KDa
Source organism: Haemophilus influenzae
Expression system: Escherichia coli
UniProt:
Pfam: Restriction endonuclease HincII
InterPro:
Length: 317 amino acids
Theoretical weight: 36.59 KDa
Source organism: Haemophilus influenzae
Expression system: Escherichia coli
UniProt:
- Canonical:
 P17743 (Residues: 1-258; Coverage: 100%)
P17743 (Residues: 1-258; Coverage: 100%)
Pfam: Restriction endonuclease HincII
InterPro:
- Restriction endonuclease type II-like
- DNA mismatch repair MutH/Type II restriction endonuclease superfamily
- Restriction endonuclease, type II, HincII
Name:
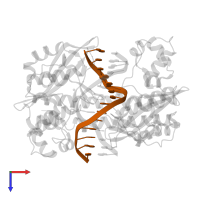
5'-D(*DGP*DCP*DCP*DGP*DGP*DTP*DCP*DGP*DAP*DCP*DGP*DGP*DGP*DC)-3'
Representative chains: E
Source organism: Haemophilus influenzae [727]
Expression system: Not provided
Length: 14 nucleotides
Theoretical weight: 4.32 KDa
Representative chains: E
Source organism: Haemophilus influenzae [727]
Expression system: Not provided
Length: 14 nucleotides
Theoretical weight: 4.32 KDa
Name:
5'-D(*DGP*DCP*DCP*DCP*DGP*DTP*DCP*DGP*DAP*DCP*DCP*DGP*DGP*DC)-3'
Representative chains: F
Source organism: Haemophilus influenzae [727]
Expression system: Not provided
Length: 14 nucleotides
Theoretical weight: 4.24 KDa
Representative chains: F
Source organism: Haemophilus influenzae [727]
Expression system: Not provided
Length: 14 nucleotides
Theoretical weight: 4.24 KDa