Assemblies
Assembly Name:
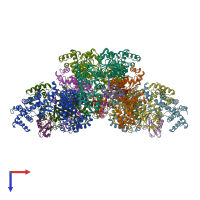
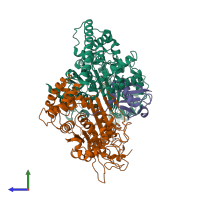
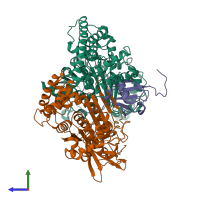
NEDD8 E1 activating enzyme complex, NAE1-UBA3 and NEDD8
Multimeric state:
hetero trimer
Accessible surface area:
41673.37 Å2
Buried surface area:
10416.34 Å2
Dissociation area:
1,707.66
Å2
Dissociation energy (ΔGdiss):
13.3
kcal/mol
Dissociation entropy (TΔSdiss):
11.8
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-171762
Assembly Name:
NEDD8 E1 activating enzyme complex, NAE1-UBA3 and NEDD8
Multimeric state:
hetero trimer
Accessible surface area:
41625.91 Å2
Buried surface area:
10525.64 Å2
Dissociation area:
1,731.47
Å2
Dissociation energy (ΔGdiss):
12.47
kcal/mol
Dissociation entropy (TΔSdiss):
11.8
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-171762
Assembly Name:
NEDD8 E1 activating enzyme complex, NAE1-UBA3 and NEDD8
Multimeric state:
hetero trimer
Accessible surface area:
41799.65 Å2
Buried surface area:
10441.62 Å2
Dissociation area:
1,686.23
Å2
Dissociation energy (ΔGdiss):
10.64
kcal/mol
Dissociation entropy (TΔSdiss):
11.79
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-171762
Assembly Name:
NEDD8 E1 activating enzyme complex, NAE1-UBA3 and NEDD8
Multimeric state:
hetero trimer
Accessible surface area:
42558.31 Å2
Buried surface area:
10373.08 Å2
Dissociation area:
1,676.79
Å2
Dissociation energy (ΔGdiss):
9.4
kcal/mol
Dissociation entropy (TΔSdiss):
12.1
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-171762
Macromolecules
Chains: A, C, E, G
Length: 531 amino acids
Theoretical weight: 59.97 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: ThiF family
InterPro:
SCOP: Ubiquitin activating enzymes (UBA)
Length: 531 amino acids
Theoretical weight: 59.97 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q13564 (Residues: 1-534; Coverage: 99%)
Q13564 (Residues: 1-534; Coverage: 99%)
Pfam: ThiF family
InterPro:
- NEDD8-activating enzyme E1 regulatory subunit APP-BP1
- Ubiquitin-activating enzyme
- ThiF/MoeB/HesA family
- THIF-type NAD/FAD binding fold
SCOP: Ubiquitin activating enzymes (UBA)
Chains: B, D, F, H
Length: 434 amino acids
Theoretical weight: 48.77 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam:
InterPro:
Length: 434 amino acids
Theoretical weight: 48.77 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q8TBC4 (Residues: 33-463; Coverage: 93%)
Q8TBC4 (Residues: 33-463; Coverage: 93%)
Pfam:
InterPro:
- Ubiquitin-activating enzyme
- ThiF/MoeB/HesA family
- THIF-type NAD/FAD binding fold
- NEDD8-activating enzyme E1 catalytic subunit, N-terminal domain
- Ubiquitin activating enzyme, alpha domain superfamily
- E2 binding
- NAD(P)-binding Rossmann-like Domain
- Ubiquitin activating enzymes (Uba3). Chain: B, domain 2
- NEDD8-activating enzyme E1, catalytic subunit
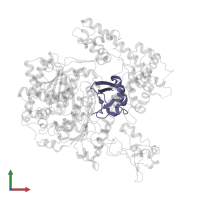
Chains: I, J, K, L
Length: 88 amino acids
Theoretical weight: 9.69 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Ubiquitin family
InterPro:
SCOP: Ubiquitin
Length: 88 amino acids
Theoretical weight: 9.69 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 Q15843 (Residues: 1-76; Coverage: 94%)
Q15843 (Residues: 1-76; Coverage: 94%)
Pfam: Ubiquitin family
InterPro:
- Ubiquitin-like domain superfamily
- Ubiquitin-like domain
- Nedd8-like ubiquitin
- Ubiquitin domain
- Ubiquitin conserved site
SCOP: Ubiquitin