Assemblies
Assembly Name:
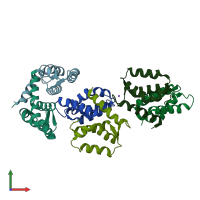
Protein S100-A12
Multimeric state:
homo dimer
Accessible surface area:
8977.11 Å2
Buried surface area:
2968.3 Å2
Dissociation area:
1,415.11
Å2
Dissociation energy (ΔGdiss):
18.18
kcal/mol
Dissociation entropy (TΔSdiss):
11.17
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-160420
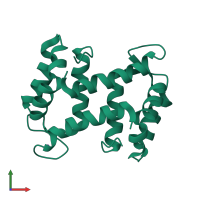
Assembly Name:
Protein S100-A12
Multimeric state:
homo dimer
Accessible surface area:
9128.16 Å2
Buried surface area:
2689.59 Å2
Dissociation area:
1,344.79
Å2
Dissociation energy (ΔGdiss):
21.67
kcal/mol
Dissociation entropy (TΔSdiss):
11.07
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-160420
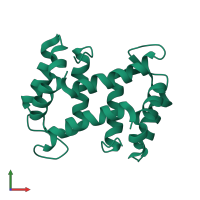
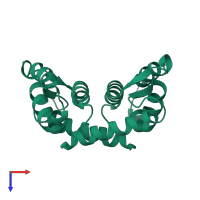
Assembly Name:
Protein S100-A12
Multimeric state:
homo dimer
Accessible surface area:
8605.25 Å2
Buried surface area:
2852.97 Å2
Dissociation area:
1,358.57
Å2
Dissociation energy (ΔGdiss):
18.64
kcal/mol
Dissociation entropy (TΔSdiss):
11.03
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-160420
Macromolecules
Chains: A, B, C, D, E, F
Length: 95 amino acids
Theoretical weight: 10.79 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: S-100/ICaBP type calcium binding domain
InterPro:
Length: 95 amino acids
Theoretical weight: 10.79 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P80511 (Residues: 2-92; Coverage: 99%)
P80511 (Residues: 2-92; Coverage: 99%)
Pfam: S-100/ICaBP type calcium binding domain
InterPro:
- EF-hand domain pair
- S100/CaBP-9k-type, calcium binding, subdomain
- EF-hand domain
- S100/Calcium binding protein 7/8-like, conserved site