Assemblies
Assembly Name:
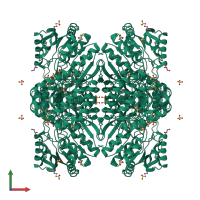
Succinate-semialdehyde dehydrogenase, mitochondrial
Multimeric state:
homo dodecamer
Accessible surface area:
194098.76 Å2
Buried surface area:
80836.55 Å2
Dissociation area:
16,376.59
Å2
Dissociation energy (ΔGdiss):
218.97
kcal/mol
Dissociation entropy (TΔSdiss):
54.26
kcal/mol
Symmetry number:
12
PDBe Complex ID:
PDB-CPX-156381
Assembly Name:
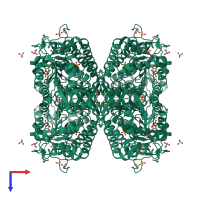
Succinate-semialdehyde dehydrogenase, mitochondrial
Multimeric state:
homo tetramer
Accessible surface area:
63491.92 Å2
Buried surface area:
28150.36 Å2
Dissociation area:
3,807.17
Å2
Dissociation energy (ΔGdiss):
59.18
kcal/mol
Dissociation entropy (TΔSdiss):
16.45
kcal/mol
Symmetry number:
4
PDBe Complex ID:
PDB-CPX-156383
Assembly Name:
Succinate-semialdehyde dehydrogenase, mitochondrial
Multimeric state:
homo dimer
Accessible surface area:
35755.63 Å2
Buried surface area:
10081.17 Å2
Dissociation area:
2,617.83
Å2
Dissociation energy (ΔGdiss):
29.07
kcal/mol
Dissociation entropy (TΔSdiss):
15.03
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-156382
Macromolecules
Chain: A
Length: 487 amino acids
Theoretical weight: 52.26 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
Pfam: Aldehyde dehydrogenase family
InterPro:
Length: 487 amino acids
Theoretical weight: 52.26 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli
UniProt:
- Canonical:
 P51649 (Residues: 49-535; Coverage: 91%)
P51649 (Residues: 49-535; Coverage: 91%)
Pfam: Aldehyde dehydrogenase family
InterPro:
- Aldehyde/histidinol dehydrogenase
- Aldehyde dehydrogenase domain
- Aldehyde dehydrogenase, N-terminal
- Succinate semialdehyde dehydrogenase
- Aldehyde dehydrogenase, glutamic acid active site
- Aldehyde dehydrogenase, C-terminal