Assemblies
Assembly Name:
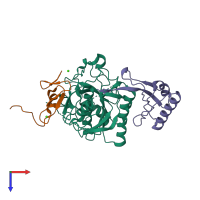
LDLR-PCSK9 complex
Multimeric state:
hetero trimer
Accessible surface area:
18295.78 Å2
Buried surface area:
3765.99 Å2
Dissociation area:
489.12
Å2
Dissociation energy (ΔGdiss):
-0.4
kcal/mol
Dissociation entropy (TΔSdiss):
10.45
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-134047
Macromolecules
Chain: A
Length: 312 amino acids
Theoretical weight: 33.11 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam: Subtilase family
InterPro:
Length: 312 amino acids
Theoretical weight: 33.11 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 Q8NBP7 (Residues: 153-451; Coverage: 45%)
Q8NBP7 (Residues: 153-451; Coverage: 45%)
Pfam: Subtilase family
InterPro:
- Peptidase S8/S53 domain superfamily
- Proteinase K-like catalytic domain
- Peptidase S8, subtilisin-related
- Peptidase S8/S53 domain
Chain: E
Length: 107 amino acids
Theoretical weight: 12.1 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
InterPro:
Length: 107 amino acids
Theoretical weight: 12.1 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 P01130 (Residues: 314-393; Coverage: 10%)
P01130 (Residues: 314-393; Coverage: 10%)
InterPro:
- Growth factor receptor cysteine-rich domain superfamily
- EGF-like domain
- EGF-like calcium-binding domain
- EGF-type aspartate/asparagine hydroxylation site
- EGF-like calcium-binding, conserved site
Chain: P
Length: 114 amino acids
Theoretical weight: 12.68 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam: Peptidase inhibitor I9
InterPro:
Length: 114 amino acids
Theoretical weight: 12.68 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 Q8NBP7 (Residues: 53-152; Coverage: 15%)
Q8NBP7 (Residues: 53-152; Coverage: 15%)
Pfam: Peptidase inhibitor I9
InterPro:
- Peptidase S8 propeptide/proteinase inhibitor I9 superfamily
- Peptidase S8 propeptide/proteinase inhibitor I9