Assemblies
Assembly Name:
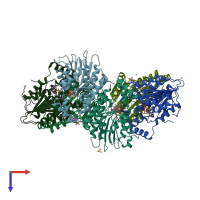
Sepiapterin reductase
Multimeric state:
homo dimer
Accessible surface area:
21375.41 Å2
Buried surface area:
7019.41 Å2
Dissociation area:
1,919.1
Å2
Dissociation energy (ΔGdiss):
31.44
kcal/mol
Dissociation entropy (TΔSdiss):
13.37
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-152880
Assembly Name:
Sepiapterin reductase
Multimeric state:
homo dimer
Accessible surface area:
20972.86 Å2
Buried surface area:
7058.88 Å2
Dissociation area:
1,917.52
Å2
Dissociation energy (ΔGdiss):
32.93
kcal/mol
Dissociation entropy (TΔSdiss):
13.35
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-152880
Assembly Name:
Sepiapterin reductase
Multimeric state:
homo dimer
Accessible surface area:
20136.25 Å2
Buried surface area:
6796.33 Å2
Dissociation area:
1,878.75
Å2
Dissociation energy (ΔGdiss):
33.46
kcal/mol
Dissociation entropy (TΔSdiss):
13.27
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-152880
Macromolecules
Chains: A, B, C, D, E, F
Length: 282 amino acids
Theoretical weight: 30.55 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam: short chain dehydrogenase
InterPro:
SCOP: Tyrosine-dependent oxidoreductases
Length: 282 amino acids
Theoretical weight: 30.55 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 P35270 (Residues: 5-261; Coverage: 99%)
P35270 (Residues: 5-261; Coverage: 99%)
Pfam: short chain dehydrogenase
InterPro:
- ATP12, ATP synthase F1-assembly protein, N-terminal
- NAD(P)-binding domain superfamily
- Sepiapterin reductase
- Short-chain dehydrogenase/reductase SDR
SCOP: Tyrosine-dependent oxidoreductases