Assemblies
Assembly Name:
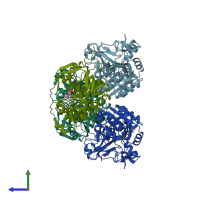
Sorbitol dehydrogenase
Multimeric state:
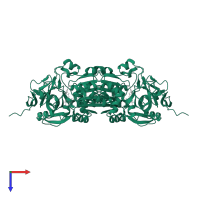
homo tetramer
Accessible surface area:
50678.01 Å2
Buried surface area:
14961.59 Å2
Dissociation area:
2,122.85
Å2
Dissociation energy (ΔGdiss):
17.96
kcal/mol
Dissociation entropy (TΔSdiss):
15.57
kcal/mol
Symmetry number:
4
PDBe Complex ID:
PDB-CPX-169474
Assembly Name:
Sorbitol dehydrogenase
Multimeric state:
homo dimer
Accessible surface area:
29157.79 Å2
Buried surface area:
3593.19 Å2
Dissociation area:
610.27
Å2
Dissociation energy (ΔGdiss):
-1.78
kcal/mol
Dissociation entropy (TΔSdiss):
13.69
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-169473
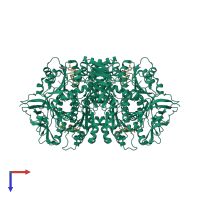
Assembly Name:
Sorbitol dehydrogenase
Multimeric state:
homo tetramer
Accessible surface area:
57446.62 Å2
Buried surface area:
8124.04 Å2
Dissociation area:
1,676.44
Å2
Dissociation energy (ΔGdiss):
-16.12
kcal/mol
Dissociation entropy (TΔSdiss):
42.07
kcal/mol
Symmetry number:
2
PDBe Complex ID:
PDB-CPX-169474
Macromolecules
Chains: A, B, C, D
Length: 356 amino acids
Theoretical weight: 38.23 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
Pfam:
InterPro:
SCOP:
Length: 356 amino acids
Theoretical weight: 38.23 KDa
Source organism: Homo sapiens
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 Q00796 (Residues: 2-357; Coverage: 100%)
Q00796 (Residues: 2-357; Coverage: 100%)
Pfam:
InterPro:
- GroES-like superfamily
- Sorbitol dehydrogenase-like
- Polyketide synthase, enoylreductase domain
- Alcohol dehydrogenase-like, N-terminal
- Alcohol dehydrogenase, zinc-type, conserved site
- NAD(P)-binding domain superfamily
- Alcohol dehydrogenase-like, C-terminal
SCOP: