Assemblies
Assembly Name:
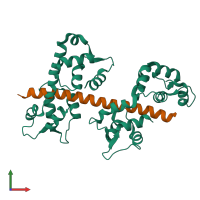
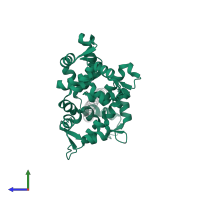
Myosin light chain 1 and Myosin-2
Multimeric state:
hetero trimer
Accessible surface area:
18799.24 Å2
Buried surface area:
5203.49 Å2
Dissociation area:
1,418.37
Å2
Dissociation energy (ΔGdiss):
10.28
kcal/mol
Dissociation entropy (TΔSdiss):
12.29
kcal/mol
Symmetry number:
1
PDBe Complex ID:
PDB-CPX-148626
Macromolecules
Chains: A, B
Length: 148 amino acids
Theoretical weight: 16.33 KDa
Source organism: Saccharomyces cerevisiae
Expression system: Escherichia coli BL21(DE3)
UniProt:
InterPro:
CATH: EF-hand
SCOP: Calmodulin-like
Length: 148 amino acids
Theoretical weight: 16.33 KDa
Source organism: Saccharomyces cerevisiae
Expression system: Escherichia coli BL21(DE3)
UniProt:
- Canonical:
 P53141 (Residues: 2-149; Coverage: 99%)
P53141 (Residues: 2-149; Coverage: 99%)
InterPro:
CATH: EF-hand
SCOP: Calmodulin-like
Chain: C
Length: 48 amino acids
Theoretical weight: 5.54 KDa
Source organism: Saccharomyces cerevisiae S288C
Expression system: Not provided
UniProt:
InterPro: IQ motif, EF-hand binding site
CATH: Single alpha-helices involved in coiled-coils or other helix-helix interfaces
Length: 48 amino acids
Theoretical weight: 5.54 KDa
Source organism: Saccharomyces cerevisiae S288C
Expression system: Not provided
UniProt:
- Canonical:
 P19524 (Residues: 806-853; Coverage: 3%)
P19524 (Residues: 806-853; Coverage: 3%)
InterPro: IQ motif, EF-hand binding site
CATH: Single alpha-helices involved in coiled-coils or other helix-helix interfaces