GO terms
Biochemical function:
Biological process:
Cellular component:
Sequence families
 Protein families (Pfam)
Protein families (Pfam)
 InterPro annotations
InterPro annotations
Structure domains
 CATH domains
CATH domains
3.40.50.300
Class:
Alpha Beta
Architecture:
3-Layer(aba) Sandwich
Topology:
Rossmann fold
Homology:
P-loop containing nucleotide triphosphate hydrolases
Occurring in:
![The deposited structure of PDB entry <span class='highlight'>1gl9</span> contains 4 copies of CATH domain <span class='highlight'>3.40.50.300 (Rossmann fold)</span> in <span class='highlight'>Reverse gyrase</span>. Showing 2 copies in chain <span class='highlight'>A [auth B]</span>. The deposited structure of PDB entry 1gl9 contains 4 copies of CATH domain 3.40.50.300 (Rossmann fold) in Reverse gyrase. Showing 2 copies in chain A [auth B].](https://www.ebi.ac.uk/pdbe/static/entry/1gl9_1_B_CATH_3.40.50.300_image-200x200.png)
2.60.510.20
Class:
Mainly Beta
Architecture:
Sandwich
Topology:
EV matrix protein fold
Homology:
EV matrix protein fold
Occurring in:
![The deposited structure of PDB entry <span class='highlight'>1gl9</span> contains 2 copies of CATH domain <span class='highlight'>2.60.510.20 (EV matrix protein fold)</span> in <span class='highlight'>Reverse gyrase</span>. Showing 1 copy in chain <span class='highlight'>A [auth B]</span>. The deposited structure of PDB entry 1gl9 contains 2 copies of CATH domain 2.60.510.20 (EV matrix protein fold) in Reverse gyrase. Showing 1 copy in chain A [auth B].](https://www.ebi.ac.uk/pdbe/static/entry/1gl9_1_B_CATH_2.60.510.20_image-200x200.png)
1.10.460.10
Class:
Mainly Alpha
Architecture:
Orthogonal Bundle
Topology:
Topoisomerase I; domain 2
Homology:
Topoisomerase I, domain 2
Occurring in:
![The deposited structure of PDB entry <span class='highlight'>1gl9</span> contains 2 copies of CATH domain <span class='highlight'>1.10.460.10 (Topoisomerase I; domain 2)</span> in <span class='highlight'>Reverse gyrase</span>. Showing 1 copy in chain <span class='highlight'>A [auth B]</span>. The deposited structure of PDB entry 1gl9 contains 2 copies of CATH domain 1.10.460.10 (Topoisomerase I; domain 2) in Reverse gyrase. Showing 1 copy in chain A [auth B].](https://www.ebi.ac.uk/pdbe/static/entry/1gl9_1_B_CATH_1.10.460.10_image-200x200.png)
3.40.50.140
Class:
Alpha Beta
Architecture:
3-Layer(aba) Sandwich
Topology:
Rossmann fold
Homology:
Rossmann fold
Occurring in:
![The deposited structure of PDB entry <span class='highlight'>1gl9</span> contains 2 copies of CATH domain <span class='highlight'>3.40.50.140 (Rossmann fold)</span> in <span class='highlight'>Reverse gyrase</span>. Showing 1 copy in chain <span class='highlight'>A [auth B]</span>. The deposited structure of PDB entry 1gl9 contains 2 copies of CATH domain 3.40.50.140 (Rossmann fold) in Reverse gyrase. Showing 1 copy in chain A [auth B].](https://www.ebi.ac.uk/pdbe/static/entry/1gl9_1_B_CATH_3.40.50.140_image-200x200.png)
2.20.20.30
Class:
Mainly Beta
Architecture:
Single Sheet
Topology:
Anthopleurin-A
Homology:
reverse gyrase domain
Occurring in:
![The deposited structure of PDB entry <span class='highlight'>1gl9</span> contains 2 copies of CATH domain <span class='highlight'>2.20.20.30 (Anthopleurin-A)</span> in <span class='highlight'>Reverse gyrase</span>. Showing 1 copy in chain <span class='highlight'>A [auth B]</span>. The deposited structure of PDB entry 1gl9 contains 2 copies of CATH domain 2.20.20.30 (Anthopleurin-A) in Reverse gyrase. Showing 1 copy in chain A [auth B].](https://www.ebi.ac.uk/pdbe/static/entry/1gl9_1_B_CATH_2.20.20.30_image-200x200.png)
1.10.290.10
Class:
Mainly Alpha
Architecture:
Orthogonal Bundle
Topology:
Topoisomerase I; domain 4
Homology:
Topoisomerase I, domain 4
Occurring in:
![The deposited structure of PDB entry <span class='highlight'>1gl9</span> contains 2 copies of CATH domain <span class='highlight'>1.10.290.10 (Topoisomerase I; domain 4)</span> in <span class='highlight'>Reverse gyrase</span>. Showing 1 copy in chain <span class='highlight'>A [auth B]</span>. The deposited structure of PDB entry 1gl9 contains 2 copies of CATH domain 1.10.290.10 (Topoisomerase I; domain 4) in Reverse gyrase. Showing 1 copy in chain A [auth B].](https://www.ebi.ac.uk/pdbe/static/entry/1gl9_1_B_CATH_1.10.290.10_image-200x200.png)
 SCOP annotations
SCOP annotations
69496
Class:
Alpha and beta proteins (a/b)
Fold:
P-loop containing nucleoside triphosphate hydrolases
Superfamily:
P-loop containing nucleoside triphosphate hydrolases
Occurring in:
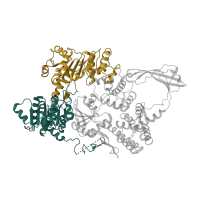 in Reverse gyrase. Showing 2 copies in chain A [auth B].">
in Reverse gyrase. Showing 2 copies in chain A [auth B].">
56713
Class:
Multi-domain proteins (alpha and beta)
Fold:
Prokaryotic type I DNA topoisomerase
Superfamily:
Prokaryotic type I DNA topoisomerase
Occurring in:
![The deposited structure of PDB entry <span class='highlight'>1gl9</span> contains 2 copies of SCOP domain <span class='highlight'>56713 (Prokaryotic type I DNA topoisomerase)</span> in <span class='highlight'>Reverse gyrase</span>. Showing 1 copy in chain <span class='highlight'>A [auth B]</span>. The deposited structure of PDB entry 1gl9 contains 2 copies of SCOP domain 56713 (Prokaryotic type I DNA topoisomerase) in Reverse gyrase. Showing 1 copy in chain A [auth B].](https://www.ebi.ac.uk/pdbe/static/entry/1gl9_1_B_SCOP_56713_image-200x200.png)



![The deposited structure of PDB entry <span class='highlight'>1gl9</span> contains 4 copies of Pfam domain <span class='highlight'>PF01131 (DNA topoisomerase)</span> in <span class='highlight'>Reverse gyrase</span>. Showing 2 copies in chain <span class='highlight'>A [auth B]</span>. The deposited structure of PDB entry 1gl9 contains 4 copies of Pfam domain PF01131 (DNA topoisomerase) in Reverse gyrase. Showing 2 copies in chain A [auth B].](https://www.ebi.ac.uk/pdbe/static/entry/1gl9_1_B_Pfam_PF01131_image-200x200.png)
![The deposited structure of PDB entry <span class='highlight'>1gl9</span> contains 2 copies of Pfam domain <span class='highlight'>PF01751 (Toprim domain)</span> in <span class='highlight'>Reverse gyrase</span>. Showing 1 copy in chain <span class='highlight'>A [auth B]</span>. The deposited structure of PDB entry 1gl9 contains 2 copies of Pfam domain PF01751 (Toprim domain) in Reverse gyrase. Showing 1 copy in chain A [auth B].](https://www.ebi.ac.uk/pdbe/static/entry/1gl9_1_B_Pfam_PF01751_image-200x200.png)
![The deposited structure of PDB entry <span class='highlight'>1gl9</span> contains 2 copies of Pfam domain <span class='highlight'>PF00270 (DEAD/DEAH box helicase)</span> in <span class='highlight'>Reverse gyrase</span>. Showing 1 copy in chain <span class='highlight'>A [auth B]</span>. The deposited structure of PDB entry 1gl9 contains 2 copies of Pfam domain PF00270 (DEAD/DEAH box helicase) in Reverse gyrase. Showing 1 copy in chain A [auth B].](https://www.ebi.ac.uk/pdbe/static/entry/1gl9_1_B_Pfam_PF00270_image-200x200.png)
![The deposited structure of PDB entry <span class='highlight'>1gl9</span> contains 2 copies of CATH domain <span class='highlight'>3.30.56.120 (Phenylalanyl-tRNA Synthetase; Chain B, domain 1)</span> in <span class='highlight'>Reverse gyrase</span>. Showing 1 copy in chain <span class='highlight'>A [auth B]</span>. The deposited structure of PDB entry 1gl9 contains 2 copies of CATH domain 3.30.56.120 (Phenylalanyl-tRNA Synthetase; Chain B, domain 1) in Reverse gyrase. Showing 1 copy in chain A [auth B].](https://www.ebi.ac.uk/pdbe/static/entry/1gl9_1_B_CATH_3.30.56.120_image-200x200.png)